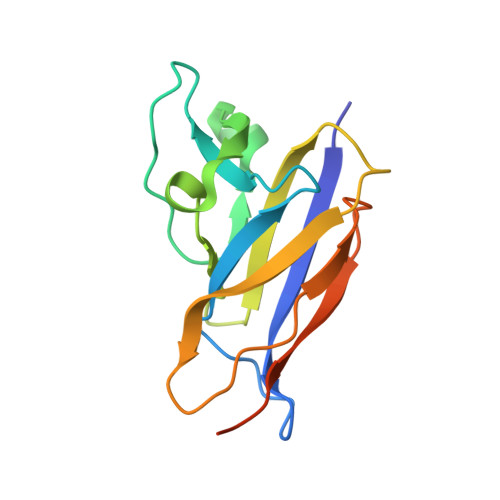
New insights about pilus formation in gut-adapted Lactobacillus rhamnosus GG from the crystal structure of the SpaA backbone-pilin subunit
Chaurasia, P., Pratap, S., von Ossowski, I., Palva, A., Krishnan, V.(2016) Sci Rep 6: 28664-28664
- PubMed: 27349405 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep28664
- Primary Citation Related Structures:
5F44, 5FAA, 5FGR, 5FGS, 5FIE, 5HBB, 5HDL, 5HTS, 5J4M - PubMed Abstract:
Thus far, all solved structures of pilin-proteins comprising sortase-assembled pili are from pathogenic genera and species. Here, we present the first crystal structure of a pilin subunit (SpaA) from a non-pathogen host (Lactobacillus rhamnosus GG). SpaA consists of two tandem CnaB-type domains, each with an isopeptide bond and E-box motif. Intriguingly, while the isopeptide bond in the N-terminal domain forms between lysine and asparagine, the one in the C-terminal domain atypically involves aspartate. We also solved crystal structures of mutant proteins where residues implicated in forming isopeptide bonds were replaced. Expectedly, the E-box-substituted E139A mutant lacks an isopeptide bond in the N-terminal domain. However, the C-terminal E269A substitution gave two structures; one of both domains with their isopeptide bonds present, and another of only the N-terminal domain, but with an unformed isopeptide bond and significant conformational changes. This latter crystal structure has never been observed for any other Gram-positive pilin. Notably, the C-terminal isopeptide bond still forms in D295N-substituted SpaA, irrespective of E269 being present or absent. Although E-box mutations affect SpaA proteolytic and thermal stability, a cumulative effect perturbing normal pilus polymerization was unobserved. A model showing the polymerized arrangement of SpaA within the SpaCBA pilus is proposed.
- Regional Centre for Biotechnology, NCR Biotech Science Cluster, Faridabad-121 001, India.
Organizational Affiliation: