Antibody Response to Hypervariable Region 1 Interferes with Broadly Neutralizing Antibodies to Hepatitis C Virus.
Keck, Z.Y., Girard-Blanc, C., Wang, W., Lau, P., Zuiani, A., Rey, F.A., Krey, T., Diamond, M.S., Foung, S.K.(2016) J Virol 90: 3112-3122
- PubMed: 26739044 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.02458-15
- Primary Citation Related Structures:
5FGB, 5FGC - PubMed Abstract:
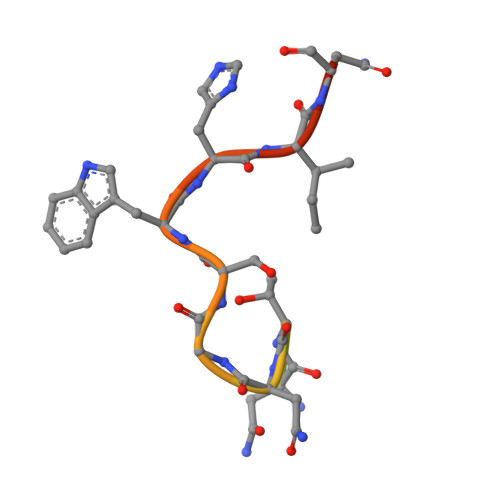
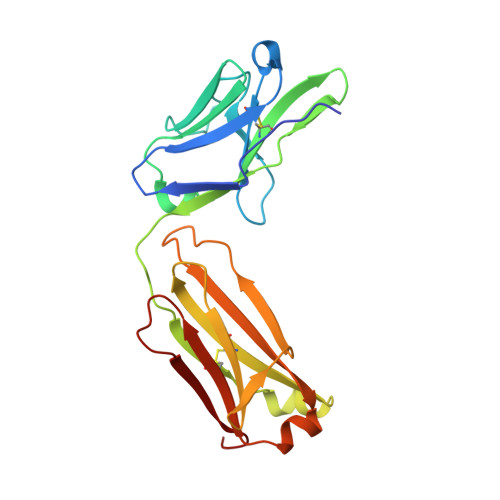
Hypervariable region 1 (HVR1) (amino acids [aa] 384 to 410) on the E2 glycoprotein of hepatitis C virus contributes to persistent infection by evolving escape mutations that attenuate binding of inhibitory antibodies and by blocking access of broadly neutralizing antibodies to their epitopes. A third proposed mechanism of immune antagonism is that poorly neutralizing antibodies binding to HVR1 interfere with binding of other superior neutralizing antibodies. Epitope mapping of human monoclonal antibodies (HMAbs) that bind to an adjacent, conserved domain on E2 encompassing aa 412 to 423 revealed two subsets, designated HC33 HMAbs. While both subsets have contact residues within aa 412 to 423, alanine-scanning mutagenesis suggested that one subset, which includes HC33.8, has an additional contact residue within HVR1. To test for interference of anti-HVR1 antibodies with binding of antibodies to aa 412 to 423 and other E2 determinants recognized by broadly neutralizing HMAbs, two murine MAbs against HVR1 (H77.16) and aa 412 to 423 (H77.39) were studied. As expected, H77.39 inhibited the binding of all HC33 HMAbs. Unexpectedly, H77.16 also inhibited the binding of both subsets of HC33 HMAbs. This inhibition also was observed against other broadly neutralizing HMAbs to epitopes outside aa 412 to 423. Combination antibody neutralization studies by the median-effect analysis method with H77.16 and broadly reactive HMAbs revealed antagonism between these antibodies. Structural studies demonstrated conformational flexibility in this antigenic region, which supports the possibility of anti-HVR1 antibodies hindering the binding of broadly neutralizing MAbs. These findings support the hypothesis that anti-HVR1 antibodies can interfere with a protective humoral response against HCV infection. HVR1 contributes to persistent infection by evolving mutations that escape from neutralizing antibodies to HVR1 and by shielding broadly neutralizing antibodies from their epitopes. This study provides insight into a new immune antagonism mechanism by which the binding of antibodies to HVR1 blocks the binding and activity of broadly neutralizing antibodies to HCV. Immunization strategies that avoid the induction of HVR1 antibodies should increase the inhibitory activity of broadly neutralizing anti-HCV antibodies elicited by candidate vaccines.
- Department of Pathology, Stanford University School of Medicine, Stanford, California, USA.
Organizational Affiliation: