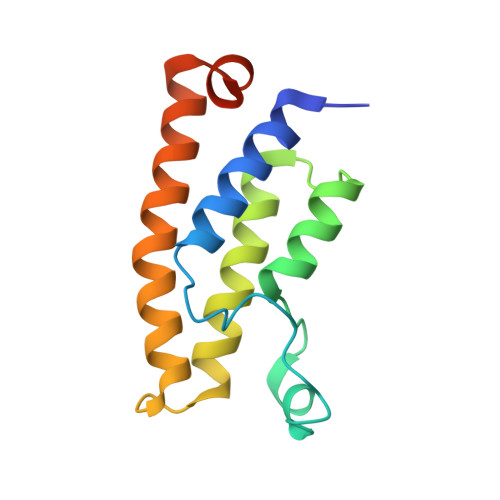
Crystal structure of the bromodomain of human BRD1 (BRPF2) in complex with OF-1 chemical probe
Tallant, C., Owen, D.R., Gerstenberger, B.S., Savitsky, P., Chaikuad, A., Fedorov, O., Nunez-Alonso, G., Filippakopoulos, P., von Delft, F., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Muller, S., Knapp, S.To be published.