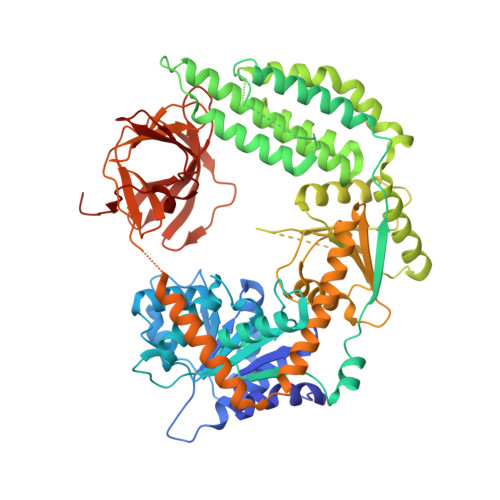
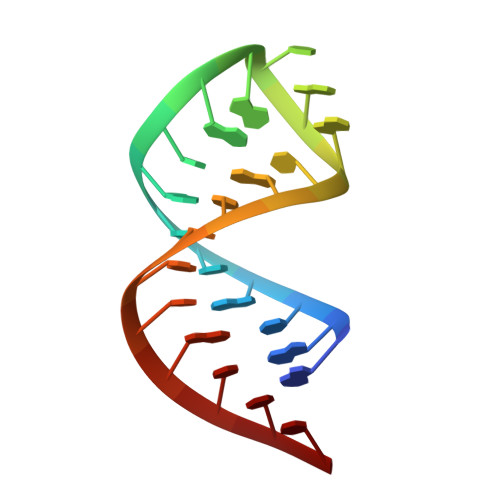
Structural basis for m7G recognition and 2'-O-methyl discrimination in capped RNAs by the innate immune receptor RIG-I.
Devarkar, S.C., Wang, C., Miller, M.T., Ramanathan, A., Jiang, F., Khan, A.G., Patel, S.S., Marcotrigiano, J.(2016) Proc Natl Acad Sci U S A 113: 596-601
- PubMed: 26733676 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1515152113
- Primary Citation Related Structures:
5F98, 5F9F, 5F9H - PubMed Abstract:
RNAs with 5'-triphosphate (ppp) are detected in the cytoplasm principally by the innate immune receptor Retinoic Acid Inducible Gene-I (RIG-I), whose activation triggers a Type I IFN response. It is thought that self RNAs like mRNAs are not recognized by RIG-I because 5'ppp is capped by the addition of a 7-methyl guanosine (m7G) (Cap-0) and a 2'-O-methyl (2'-OMe) group to the 5'-end nucleotide ribose (Cap-1). Here we provide structural and mechanistic basis for exact roles of capping and 2'-O-methylation in evading RIG-I recognition. Surprisingly, Cap-0 and 5'ppp double-stranded (ds) RNAs bind to RIG-I with nearly identical Kd values and activate RIG-I's ATPase and cellular signaling response to similar extents. On the other hand, Cap-0 and 5'ppp single-stranded RNAs did not bind RIG-I and are signaling inactive. Three crystal structures of RIG-I complexes with dsRNAs bearing 5'OH, 5'ppp, and Cap-0 show that RIG-I can accommodate the m7G cap in a cavity created through conformational changes in the helicase-motif IVa without perturbing the ppp interactions. In contrast, Cap-1 modifications abrogate RIG-I signaling through a mechanism involving the H830 residue, which we show is crucial for discriminating between Cap-0 and Cap-1 RNAs. Furthermore, m7G capping works synergistically with 2'-O-methylation to weaken RNA affinity by 200-fold and lower ATPase activity. Interestingly, a single H830A mutation restores both high-affinity binding and signaling activity with 2'-O-methylated dsRNAs. Our work provides new structural insights into the mechanisms of host and viral immune evasion from RIG-I, explaining the complexity of cap structures over evolution.
- Department of Biochemistry and Molecular Biology, Robert Wood Johnson Medical School, Rutgers University, Piscataway, NJ 08854;
Organizational Affiliation: