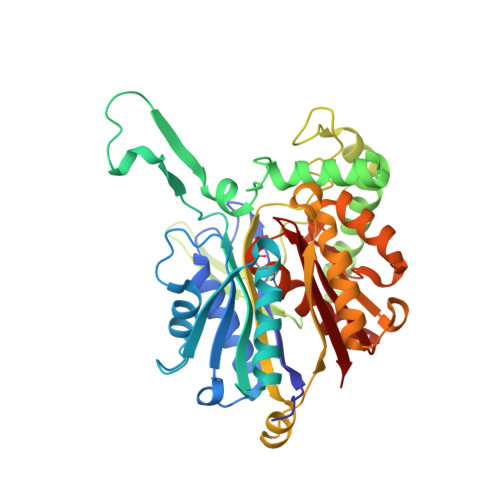
Crystal structure of a thiolase from Escherichia coli at 1.8 angstrom resolution.
Ithayaraja, M., Janardan, N., Wierenga, R.K., Savithri, H.S., Murthy, M.R.(2016) Acta Crystallogr F Struct Biol Commun 72: 534-544
- PubMed: 27380370 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X16008451
- Primary Citation Related Structures:
5F0V, 5F38 - PubMed Abstract:
Thiolases catalyze the Claisen condensation of two acetyl-CoA molecules to give acetoacetyl-CoA, as well as the reverse degradative reaction. Four genes coding for thiolases or thiolase-like proteins are found in the Escherichia coli genome. In this communication, the successful cloning, purification, crystallization and structure determination at 1.8 Å resolution of a homotetrameric E. coli thiolase are reported. The structure of E. coli thiolase co-crystallized with acetyl-CoA at 1.9 Å resolution is also reported. As observed in other tetrameric thiolases, the present E. coli thiolase is a dimer of two tight dimers and probably functions as a biodegradative enzyme. Comparison of the structure and biochemical properties of the E. coli enzyme with those of other well studied thiolases reveals certain novel features of this enzyme, such as the modification of a lysine in the dimeric interface, the possible oxidation of the catalytic Cys88 in the structure of the enzyme obtained in the presence of CoA and active-site hydration. The tetrameric enzyme also displays an interesting departure from exact 222 symmetry, which is probably related to the deformation of the tetramerization domain that stabilizes the oligomeric structure of the protein. The current study allows the identification of substrate-binding amino-acid residues and water networks at the active site and provides the structural framework required for understanding the biochemical properties as well as the physiological function of this E. coli thiolase.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, Karnataka 560 012, India.
Organizational Affiliation: