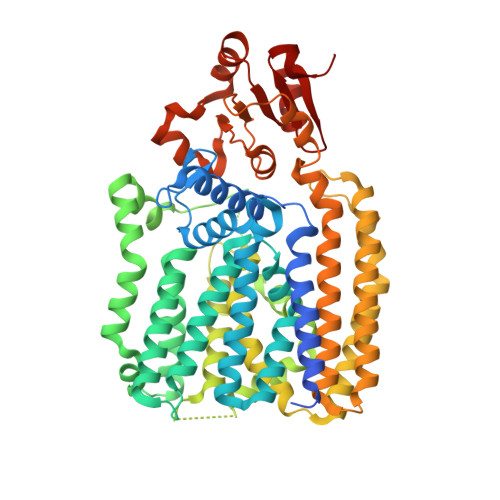
Structures of aminoarabinose transferase ArnT suggest a molecular basis for lipid A glycosylation.
Petrou, V.I., Herrera, C.M., Schultz, K.M., Clarke, O.B., Vendome, J., Tomasek, D., Banerjee, S., Rajashankar, K.R., Belcher Dufrisne, M., Kloss, B., Kloppmann, E., Rost, B., Klug, C.S., Trent, M.S., Shapiro, L., Mancia, F.(2016) Science 351: 608-612
- PubMed: 26912703 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.aad1172
- Primary Citation Related Structures:
5EZM, 5F15 - PubMed Abstract:
Polymyxins are antibiotics used in the last line of defense to combat multidrug-resistant infections by Gram-negative bacteria. Polymyxin resistance arises through charge modification of the bacterial outer membrane with the attachment of the cationic sugar 4-amino-4-deoxy-l-arabinose to lipid A, a reaction catalyzed by the integral membrane lipid-to-lipid glycosyltransferase 4-amino-4-deoxy-L-arabinose transferase (ArnT). Here, we report crystal structures of ArnT from Cupriavidus metallidurans, alone and in complex with the lipid carrier undecaprenyl phosphate, at 2.8 and 3.2 angstrom resolution, respectively. The structures show cavities for both lipidic substrates, which converge at the active site. A structural rearrangement occurs on undecaprenyl phosphate binding, which stabilizes the active site and likely allows lipid A binding. Functional mutagenesis experiments based on these structures suggest a mechanistic model for ArnT family enzymes.
- Department of Physiology and Cellular Biophysics, Columbia University, New York, NY 10032, USA.
Organizational Affiliation: