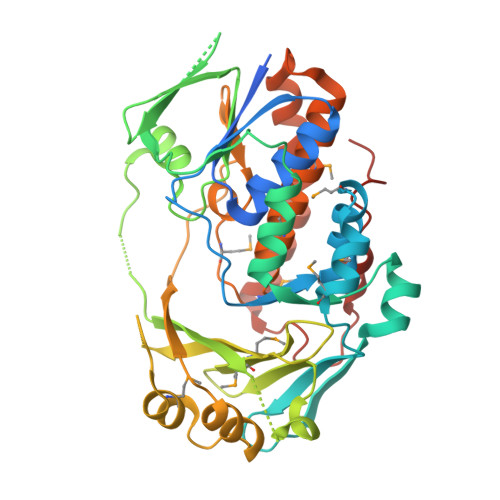
Structural and Biochemical Characterization of 6-Hydroxynicotinic Acid 3-Monooxygenase, A Novel Decarboxylative Hydroxylase Involved in Aerobic Nicotinate Degradation.
Hicks, K.A., Yuen, M.E., Zhen, W.F., Gerwig, T.J., Story, R.W., Kopp, M.C., Snider, M.J.(2016) Biochemistry 55: 3432-3446
- PubMed: 27218267 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.6b00105
- Primary Citation Related Structures:
5EOW - Department of Chemistry, SUNY Cortland , Cortland, New York 13045, United States.
Organizational Affiliation: