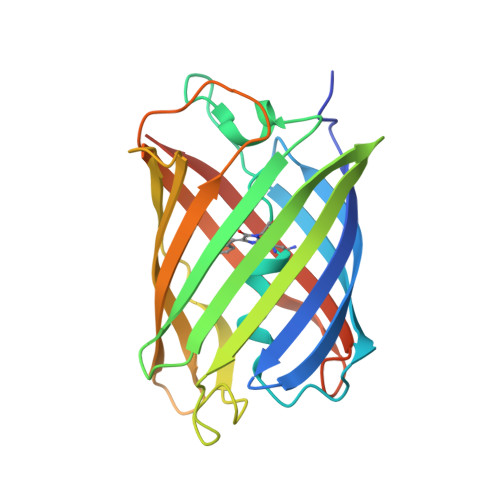
Evolution and characterization of a new reversibly photoswitching chromogenic protein, Dathail.
Langan, P.S., Close, D.W., Coates, L., Rocha, R.C., Ghosh, K., Kiss, C., Waldo, G., Freyer, J., Kovalevsky, A., Bradbury, A.R.(2016) J Mol Biology 428: 1776-1789
- PubMed: 27000644 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2016.02.029
- Primary Citation Related Structures:
5EB6, 5EB7, 5EBJ, 5EJU, 5EXU - PubMed Abstract:
We report the engineering of a new reversibly switching chromogenic protein, Dathail. Dathail was evolved from the extremely thermostable fluorescent proteins thermal green protein (TGP) and eCGP123 using directed evolution and ratiometric sorting. Dathail has two spectrally distinct chromogenic states with low quantum yields, corresponding to absorbance in a ground state with a maximum at 389nm, and a photo-induced metastable state with a maximum at 497nm. In contrast to all previously described photoswitchable proteins, both spectral states of Dathail are non-fluorescent. The photo-induced chromogenic state of Dathail has a lifetime of ~50min at 293K and pH7.5 as measured by UV-Vis spectrophotometry, returning to the ground state through thermal relaxation. X-ray crystallography provided structural insights supporting a change in conformation and coordination in the chromophore pocket as being responsible for Dathail's photoswitching. Neutron crystallography, carried out for the first time on a protein from the green fluorescent protein family, showed a distribution of hydrogen atoms revealing protonation of the chromophore 4-hydroxybenzyl group in the ground state. The neutron structure also supports the hypothesis that the photo-induced proton transfer from the chromophore occurs through water-mediated proton relay into the bulk solvent. Beyond its spectroscopic curiosity, Dathail has several characteristics that are improvements for applications, including low background fluorescence, large spectral separation, rapid switching time, and the ability to switch many times. Therefore, Dathail is likely to be extremely useful in the quickly developing fields of imaging and biosensors, including photochromic Förster resonance energy transfer, high-resolution microscopy, and live tracking within the cell.
- Bioscience Division, Los Alamos National Laboratory, Los Alamos, NM 87545, USA; Nanoscience and Microsystems Engineering, University of New Mexico, Albuquerque, NM 87131, USA.
Organizational Affiliation: