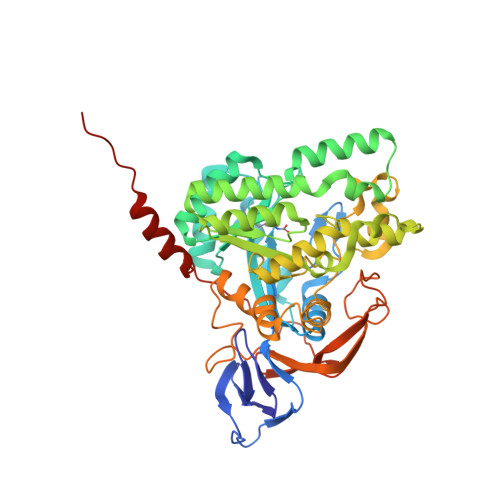
Crystal structure of dihydropyrimidinase from Pseudomonas aeruginosa PAO1: Insights into the molecular basis of formation of a dimer
Tzeng, C.T., Huang, Y.H., Huang, C.Y.(2016) Biochem Biophys Res Commun 478: 1449-1455
- PubMed: 27576201 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2016.08.144
- Primary Citation Related Structures:
5E5C - PubMed Abstract:
Dihydropyrimidinase, a tetrameric metalloenzyme, is a member of the cyclic amidohydrolase family, which also includes allantoinase, dihydroorotase, hydantoinase, and imidase. In this paper, we report the crystal structure of dihydropyrimidinase from Pseudomonas aeruginosa PAO1 at 2.1 Å resolution. The structure of P. aeruginosa dihydropyrimidinase reveals a classic (β/α)8-barrel structure core embedding the catalytic dimetal center and a β-sandwich domain, which is commonly found in the architecture of dihydropyrimidinases. In contrast to all dihydropyrimidinases, P. aeruginosa dihydropyrimidinase forms a dimer, rather than a tetramer, both in the crystalline state and in the solution. Basing on sequence analysis and structural comparison of the C-terminal region and the dimer-dimer interface between P. aeruginosa dihydropyrimidinase and Thermus sp. dihydropyrimidinase, we propose a working model to explain why this enzyme cannot be a tetramer.
- School of Biomedical Sciences, Chung Shan Medical University, No. 110, Sec. 1, Chien-Kuo N. Rd., Taichung City, Taiwan.
Organizational Affiliation: