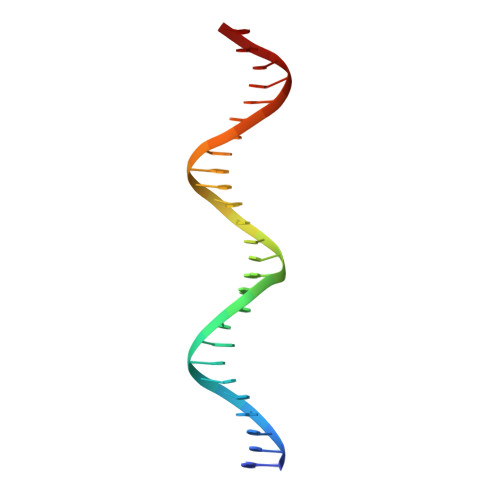
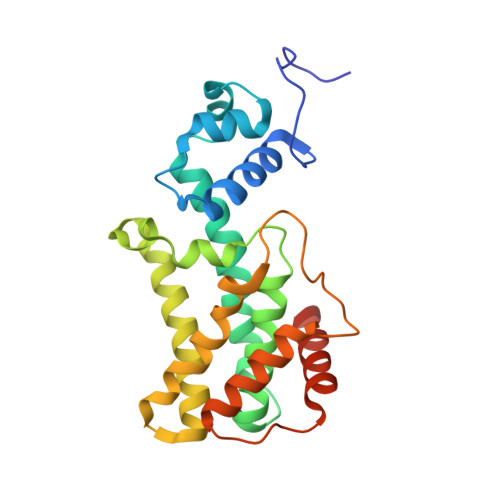
Structure of AmtR, the global nitrogen regulator of Corynebacterium glutamicum, in free and DNA-bound forms.
Palanca, C., Rubio, V.(2016) FEBS J 283: 1039-1059
- PubMed: 26744254 Search on PubMed
- DOI: https://doi.org/10.1111/febs.13643
- Primary Citation Related Structures:
5DXZ, 5DY0, 5DY1 - PubMed Abstract:
Corynebacterium glutamicum is a bacterium used for industrial amino acid production, and understanding its metabolic pathway regulation is of high biotechnological interest. Here, we report crystal structures of AmtR, the global nitrogen regulator of C. glutamicum, in apo (2.25-Å and 2.65-Å resolution) and DNA-bound (3-Å resolution) forms. These structures reveal an all-α homodimeric TetR family regulator composed of a helix-turn-helix-hosting N-terminal DNA-binding domain and a C-terminal dimerization domain. AmtR has several unique structural features that appear to be invariant among AmtR proteins, which may be related to its regulation by the nitrogen-sensing trimeric protein GlnK rather than by small-molecule effectors. As compared with other TetR family members, AmtR has an extra C-terminal helix, a large extended external loop that resembles the flexible tranducer T-loop of GlnK in sequence, and a large open cavity towards the intersubunit region that changes shape upon DNA binding. The marked kinking of helix 4 decreases in the DNA-bound form. The binding of one AmtR dimer to its DNA operator involves not only the insertion of helices 3 and 3' in adjacent turns of the double-helix major groove, but also the anchoring of 19-residue, arginine-rich and proline-rich N-terminal extensions to two external minor grooves. Electrophoretic mobility shift assays with a deletion mutant reveal that the 19-residue extension is crucial for AmtR binding to DNA. N-extension anchoring explains the flanking by AT sequences of the recognized target DNA sequence core. The significance of these findings for the entire TetR family of regulators and for GlnK regulation of AmtR is discussed. The atomic coordinates and structure factors have been deposited in the Protein Data Bank, www.pdb.org [PDB ID codes 5DXZ (native AmtR), 5DY1 (SeMet-AmtR), and 5DY0 (AmtR·DNA)].
- Instituto de Biomedicina de Valencia of the CSIC (IBV-CSIC), Spain.
Organizational Affiliation: