Crystal Structure of E. coli YbgJK
Arbing, M.A., Kaufmann, M., Shin, A., Medrano-Soto, A., Cascio, D., Eisenberg, D.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| YbgK | 310 | Escherichia coli K-12 | Mutation(s): 0 Gene Names: ybgK, b0712, JW0702 EC: 3.5.2.9 | 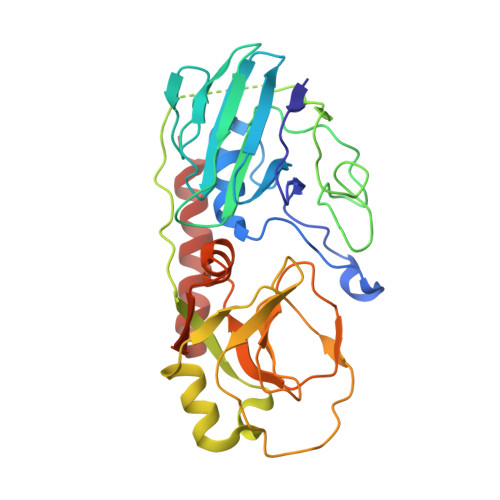 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P75745 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| YbgJ | 218 | Escherichia coli K-12 | Mutation(s): 0 Gene Names: ybgJ, b0711, JW0701 EC: 3.5.2.9 | 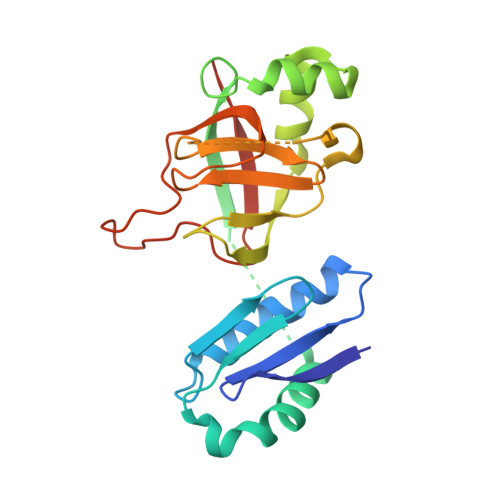 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0AAV4 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 66.11 | α = 90 |
| b = 88.78 | β = 101.91 |
| c = 83.7 | γ = 90 |
| Software Name | Purpose |
|---|---|
| BUSTER | refinement |
| XDS | data reduction |
| XSCALE | data scaling |
| MR-Rosetta | phasing |
| PDB_EXTRACT | data extraction |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | GM071909-01A2 |