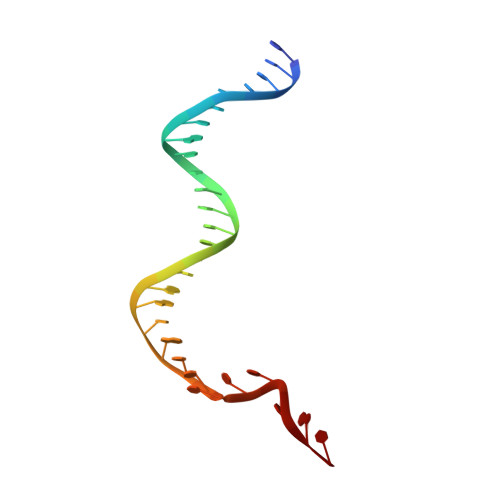
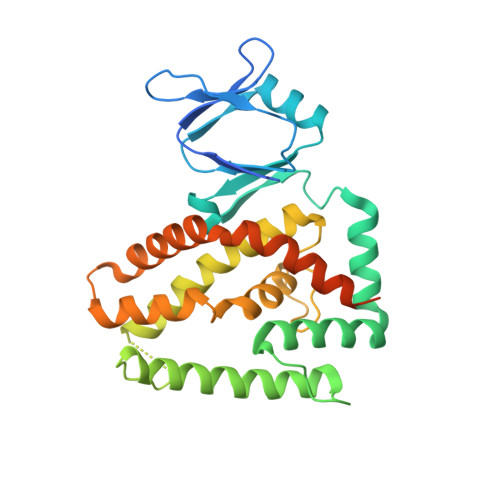
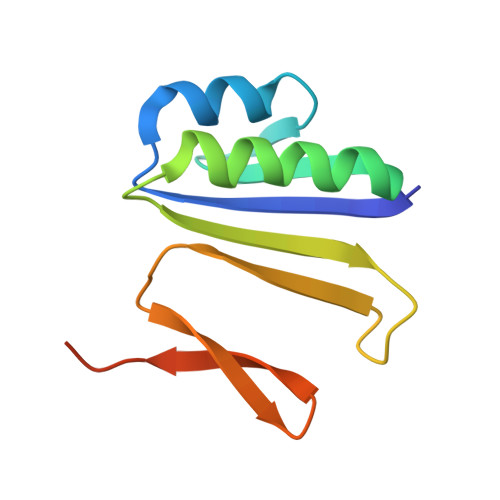
Foreign DNA capture during CRISPR-Cas adaptive immunity.
Nunez, J.K., Harrington, L.B., Kranzusch, P.J., Engelman, A.N., Doudna, J.A.(2015) Nature 527: 535-538
- PubMed: 26503043 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature15760
- Primary Citation Related Structures:
5DS4, 5DS5, 5DS6 - PubMed Abstract:
Bacteria and archaea generate adaptive immunity against phages and plasmids by integrating foreign DNA of specific 30-40-base-pair lengths into clustered regularly interspaced short palindromic repeat (CRISPR) loci as spacer segments. The universally conserved Cas1-Cas2 integrase complex catalyses spacer acquisition using a direct nucleophilic integration mechanism similar to retroviral integrases and transposases. How the Cas1-Cas2 complex selects foreign DNA substrates for integration remains unknown. Here we present X-ray crystal structures of the Escherichia coli Cas1-Cas2 complex bound to cognate 33-nucleotide protospacer DNA substrates. The protein complex creates a curved binding surface spanning the length of the DNA and splays the ends of the protospacer to allow each terminal nucleophilic 3'-OH to enter a channel leading into the Cas1 active sites. Phosphodiester backbone interactions between the protospacer and the proteins explain the sequence-nonspecific substrate selection observed in vivo. Our results uncover the structural basis for foreign DNA capture and the mechanism by which Cas1-Cas2 functions as a molecular ruler to dictate the sequence architecture of CRISPR loci.
- Department of Molecular and Cell Biology, University of California, Berkeley, Berkeley, California 94720, USA.
Organizational Affiliation: