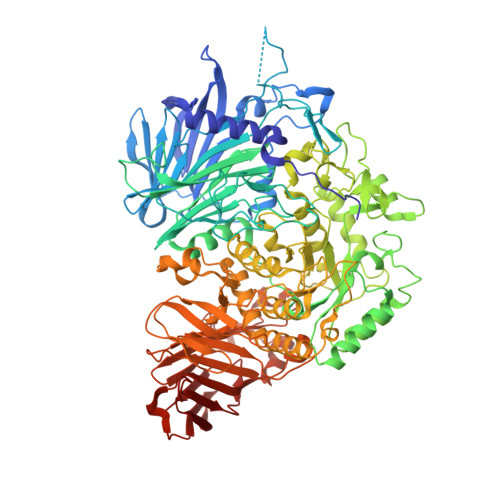
Structural basis for two-step glucose trimming by glucosidase II involved in ER glycoprotein quality control.
Satoh, T., Toshimori, T., Yan, G., Yamaguchi, T., Kato, K.(2016) Sci Rep 6: 20575-20575
- PubMed: 26847925 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep20575
- Primary Citation Related Structures:
5DKX, 5DKY, 5DKZ, 5DL0 - PubMed Abstract:
The endoplasmic reticulum (ER) has a sophisticated protein quality control system for the efficient folding of newly synthesized proteins. In this system, a variety of N-linked oligosaccharides displayed on proteins serve as signals recognized by series of intracellular lectins. Glucosidase II catalyzes two-step hydrolysis at α1,3-linked glucose-glucose and glucose-mannose residues of high-mannose-type glycans to generate a quality control protein tag that is transiently expressed on glycoproteins and recognized by ER chaperones. Here we determined the crystal structures of the catalytic α subunit of glucosidase II (GIIα) complexed with two different glucosyl ligands containing the scissile bonds of first- and second-step reactions. Our structural data revealed that the nonreducing terminal disaccharide moieties of the two kinds of substrates can be accommodated in a gourd-shaped bilocular pocket, thereby providing a structural basis for substrate-binding specificity in the two-step deglucosylation catalyzed by this enzyme.
- Graduate School of Pharmaceutical Sciences, Nagoya City University, 3-1 Tanabe-dori, Mizuho-ku, Nagoya 467-8603, Japan.
Organizational Affiliation: