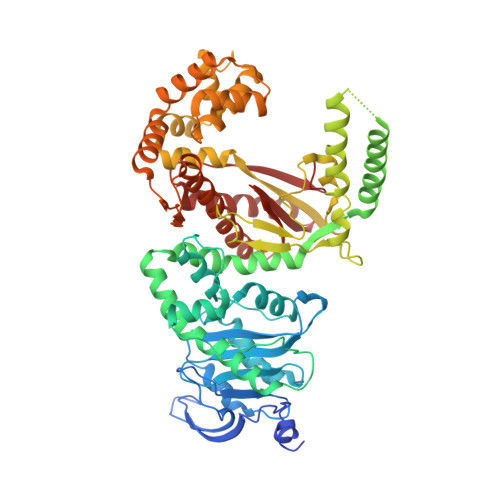
Crystal Structure of the Apicoplast DNA Polymerase from Plasmodium falciparum: The First Look at a Plastidic A-Family DNA Polymerase.
Milton, M.E., Choe, J.Y., Honzatko, R.B., Nelson, S.W.(2016) J Mol Biology 428: 3920-3934
- PubMed: 27487482 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2016.07.016
- Primary Citation Related Structures:
5DKT, 5DKU - PubMed Abstract:
Plasmodium falciparum, the primary cause of malaria, contains a non-photosynthetic plastid called the apicoplast. The apicoplast exists in most members of the phylum Apicomplexa and has its own genome along with organelle-specific enzymes for its replication. The only DNA polymerase found in the apicoplast (apPOL) was putatively acquired through horizontal gene transfer from a bacteriophage and is classified as an atypical A-family polymerase. Here, we present its crystal structure at a resolution of 2.9Å. P. falciparum apPOL, the first structural representative of a plastidic A-family polymerase, diverges from typical A-family members in two of three previously identified signature motifs and in a region not implicated by sequence. Moreover, apPOL has an additional N-terminal subdomain, the absence of which severely diminishes its 3' to 5' exonuclease activity. A compound known to be toxic to Plasmodium is a potent inhibitor of apPOL, suggesting that apPOL is a viable drug target. The structure provides new insights into the structural diversity of A-family polymerases and may facilitate structurally guided antimalarial drug design.
- Department of Biochemistry, Biophysics, and Molecular Biology, Iowa State University, Ames, IA 50011, USA.
Organizational Affiliation: