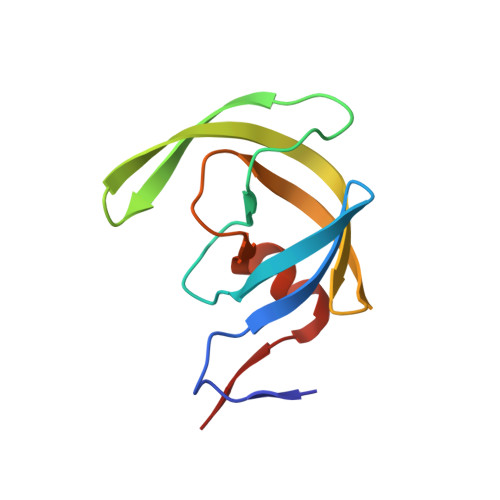Design, synthesis, biological evaluation and X-ray structural studies of HIV-1 protease inhibitors containing substituted fused-tetrahydropyranyl tetrahydrofuran as P2-ligands.
Ghosh, A.K., Martyr, C.D., Kassekert, L.A., Nyalapatla, P.R., Steffey, M., Agniswamy, J., Wang, Y.F., Weber, I.T., Amano, M., Mitsuya, H.(2015) Org Biomol Chem 13: 11607-11621
- PubMed: 26462551 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/c5ob01930c
- Primary Citation Related Structures:
5DGU, 5DGW - PubMed Abstract:
Design, synthesis, biological and X-ray crystallographic studies of a series of potent HIV-1 protease inhibitors are described. Various polar functionalities have been incorporated on the tetrahydropyranyl-tetrahydrofuran-derived P2 ligand to interact with the backbone atoms in the S2-subsite. The majority of the inhibitors showed very potent enzyme inhibitory and antiviral activity. Two high-resolution X-ray structures of 30b- and 30j-bound HIV-1 protease provide insight into ligand-binding site interactions. In particular, the polar functionalities on the P2-ligand appear to form unique hydrogen bonds with Gly48 amide NH and amide carbonyl groups in the flap region.
- Departments of Chemistry and Medicinal Chemistry, Purdue University, West Lafayette, IN 47907, USA. akghosh@purdue.edu.
Organizational Affiliation: