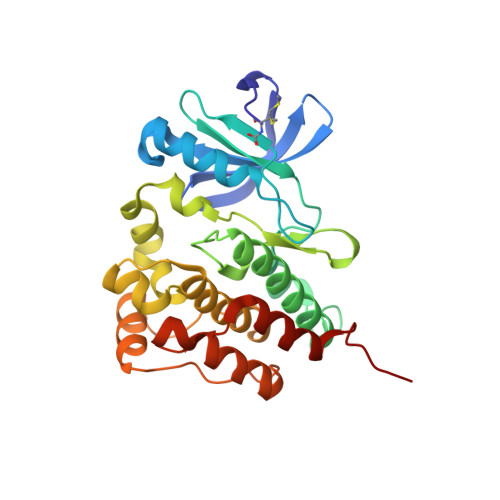
Crystal structure of the kinase domain of human protein tyrosine kinase 6 (PTK6) at 2.33 angstrom resolution
Thakur, M.K., Kumar, A., Birudukota, S., Swaminathan, S., Tyagi, R., Gosu, R.(2016) Biochem Biophys Res Commun 478: 637-642
- PubMed: 27480927 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2016.07.121
- Primary Citation Related Structures:
5D7V - PubMed Abstract:
Human Protein tyrosine kinase 6 (PTK6) (EC:2.7.10.2), also known as the breast tumor kinase (BRK), is an intracellular non-receptor Src-related tyrosine kinase expressed in a majority of human breast tumors and breast cancer cell lines, but its expression is low or completely absent in normal mammary glands. In the recent past, several studies have suggested that PTK6 is a potential therapeutic target in cancer. To understand its structural and functional properties, the PTK6 kinase domain (PTK6-KD) gene was cloned, overexpressed in a baculo-insect cell system, purified and crystallized at room temperature. X-ray diffraction data to 2.33 Å resolution was collected on a single PTK6-KD crystal, which belonged to the triclinic space group P1. The Matthews coefficient calculation suggested the presence of four protein molecules per asymmetric unit, with a solvent content of ∼50%.The structure has been solved by molecular replacement and crystal structure data submitted to the protein data bank under the accession number 5D7V. This is the first report of apo PTK6-KD structure crystallized in DFG-in and αC-helix-out conformation.
- Department of Biochemistry, University of Mysore, Mysore, 570005, India; Department of Structural Biology, Jubilant Biosys Ltd, Bangalore, 560022, India.
Organizational Affiliation: