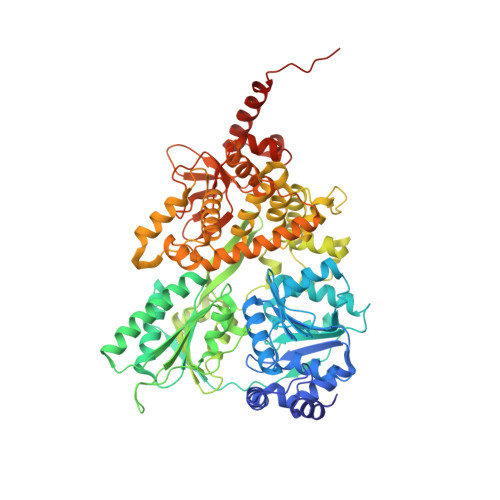
Structural and functional analysis of the RNA helicase Prp43 from the thermophilic eukaryote Chaetomium thermophilum.
Tauchert, M.J., Fourmann, J.B., Christian, H., Luhrmann, R., Ficner, R.(2016) Acta Crystallogr F Struct Biol Commun 72: 112-120
- PubMed: 26841761 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X15024498
- Primary Citation Related Structures:
5D0U - PubMed Abstract:
RNA helicases are indispensable for all organisms in each domain of life and have implications in numerous cellular processes. The DEAH-box RNA helicase Prp43 is involved in pre-mRNA splicing as well as rRNA maturation. Here, the crystal structure of Chaetomium thermophilum Prp43 at 2.9 Å resolution is revealed. Furthermore, it is demonstrated that Prp43 from C. thermophilum is capable of functionally replacing its orthologue from Saccharomyces cerevisiae in spliceosomal disassembly assays.
- Department of Molecular Structural Biology, Institute for Microbiology and Genetics, GZMB, Georg-August-Universität Göttingen, Justus-von-Liebig Weg 11, 37077 Göttingen, Germany.
Organizational Affiliation: