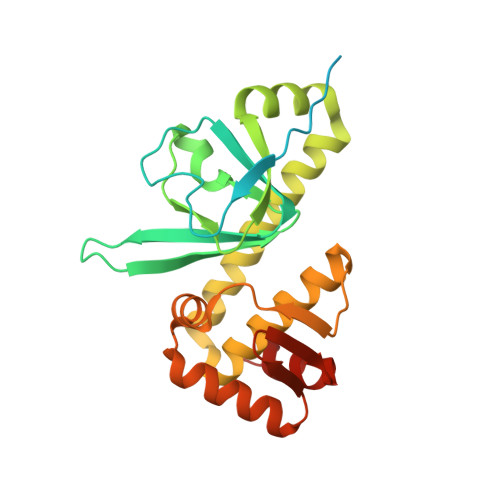
The crystal structure of the global anaerobic transcriptional regulator FNR explains its extremely fine-tuned monomer-dimer equilibrium.
Volbeda, A., Darnault, C., Renoux, O., Nicolet, Y., Fontecilla-Camps, J.C.(2015) Sci Adv 1: e1501086-e1501086
- PubMed: 26665177 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.1501086
- Primary Citation Related Structures:
5CVR, 5E44 - PubMed Abstract:
The structure of the dimeric holo-fumarate and nitrate reduction regulator (FNR) from Aliivibrio fischeri has been solved at 2.65 Å resolution. FNR globally controls the transition between anaerobic and aerobic respiration in facultative anaerobes through the assembly/degradation of its oxygen-sensitive [4Fe-4S] cluster. In the absence of O2, FNR forms a dimer and specifically binds to DNA, whereas in its presence, the cluster is degraded causing FNR monomerization and DNA dissociation. We have used our crystal structure and the information previously gathered from numerous FNR variants to propose that this process is governed by extremely fine-tuned interactions, mediated by two salt bridges near the amino-terminal cluster-binding domain and an "imperfect" coiled-coil dimer interface. [4Fe-4S] to [2Fe-2S] cluster degradation propagates a conformational signal that indirectly causes monomerization by disrupting the first of these interactions and unleashing the "unzipping" of the FNR dimer in the direction of the carboxyl-terminal DNA binding domain.
- Metalloproteins Unit, Institut de Biologie Structurale, CEA, CNRS, Université Grenoble Alpes, 71, Avenue des Martyrs CS10090, 38044 Grenoble, France.
Organizational Affiliation: