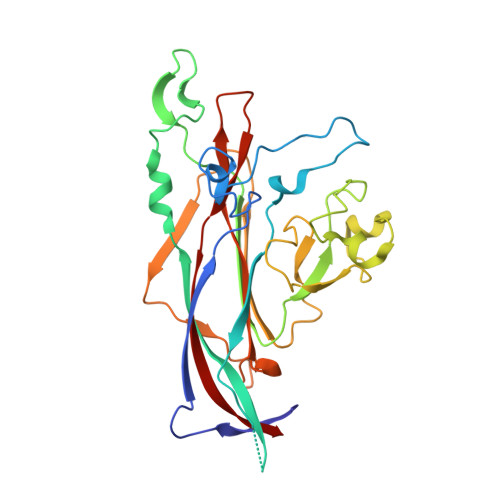
Structural and Functional Analysis of Murine Polyomavirus Capsid Proteins Establish the Determinants of Ligand Recognition and Pathogenicity.
Buch, M.H., Liaci, A.M., O'Hara, S.D., Garcea, R.L., Neu, U., Stehle, T.(2015) PLoS Pathog 11: e1005104-e1005104
- PubMed: 26474293 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1005104
- Primary Citation Related Structures:
5CPU, 5CPW, 5CPX, 5CPY, 5CPZ, 5CQ0 - PubMed Abstract:
Murine polyomavirus (MuPyV) causes tumors of various origins in newborn mice and hamsters. Infection is initiated by attachment of the virus to ganglioside receptors at the cell surface. Single amino acid exchanges in the receptor-binding pocket of the major capsid protein VP1 are known to drastically alter tumorigenicity and spread in closely related MuPyV strains. The virus represents a rare example of differential receptor recognition directly influencing viral pathogenicity, although the factors underlying these differences remain unclear. We performed structural and functional analyses of three MuPyV strains with strikingly different pathogenicities: the low-tumorigenicity strain RA, the high-pathogenicity strain PTA, and the rapidly growing, lethal laboratory isolate strain LID. Using ganglioside deficient mouse embryo fibroblasts, we show that addition of specific gangliosides restores infectability for all strains, and we uncover a complex relationship between virus attachment and infection. We identify a new infectious ganglioside receptor that carries an additional linear [α-2,8]-linked sialic acid. Crystal structures of all three strains complexed with representative oligosaccharides from the three main pathways of ganglioside biosynthesis provide the molecular basis of receptor recognition. All strains bind to a range of sialylated glycans featuring the central [α-2,3]-linked sialic acid present in the established receptors GD1a and GT1b, but the presence of additional sialic acids modulates binding. An extra [α-2,8]-linked sialic acid engages a protein pocket that is conserved among the three strains, while another, [α-2,6]-linked branching sialic acid lies near the strain-defining amino acids but can be accommodated by all strains. By comparing electron density of the oligosaccharides within the binding pockets at various concentrations, we show that the [α-2,8]-linked sialic acid increases the strength of binding. Moreover, the amino acid exchanges have subtle effects on their affinity for the validated receptor GD1a. Our results indicate that both receptor specificity and affinity influence MuPyV pathogenesis.
- Interfaculty Institute of Biochemistry, University of Tübingen, Tübingen, Germany.
Organizational Affiliation: