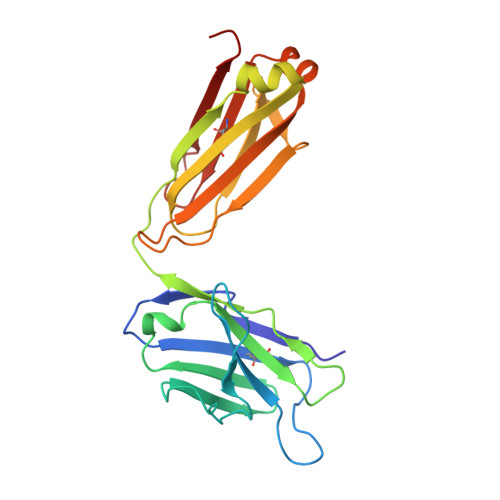
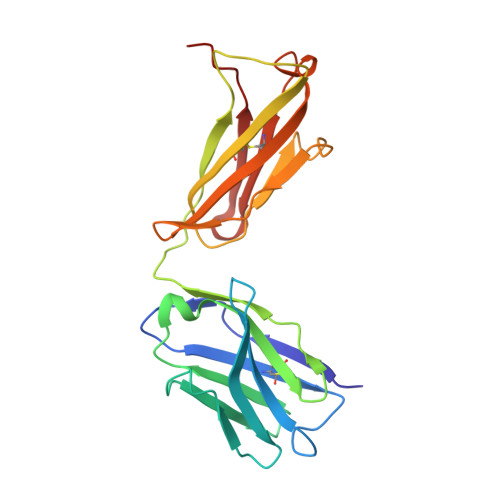
Class-specific Monoclonal Antibodies and Dihydropteroate Synthase in Bioassays used for the Detection of Sulfonamides: Structural Insights into Recognition Diversity.
Li, C., Liang, X., Wen, K., Li, Y., Zhang, X., Ma, M., Yu, X., Yu, W., Shen, J., Wang, Z.(2019) Anal Chem 91: 2392-2400
- PubMed: 30580515 Search on PubMed
- DOI: https://doi.org/10.1021/acs.analchem.8b05174
- Primary Citation Related Structures:
5CP3, 5CP7 - PubMed Abstract:
Molecular recognition between a receptor and ligand is a fundamental event in bioanalytical assays, which guarantees the sensitivity and specificity of an assay for the detection of the target of interest. An intensive understanding of the interaction mechanism could be useful for desirable hapten design, directed antibody evolution in vitro, and assay improvement. To illustrate the structural information on class-specific monoclonal antibodies (mAbs) and dihydropteroate synthase (DHPS) against sulfonamides (SAs) we have previously prepared, we initially measured the kinetic parameters of mAb 4C7, 4D11, and DHPS, which showed that the affinities of 4C7 and 4D11 were in the pM to μM range, while DHPS was uniformly in the μM range. Three-dimensional quantitative structure-activity relationship analysis for 4C7 and 4D11 then revealed that the contributions from the stereochemical structure and electron density of the SAs were comparable to binding with mAb. To acquire insights into the structural basis of mAbs and DHPS during the recognition process, the crystal structures of 4C7 and its complex with sulfathiazole were determined using X-ray crystallography. The results showed the SAs orientation and hydrogen bonding interactions mainly contributed to the diverse SAs-mAb affinities. However, for DHPS, a nucleophilic substitution reaction occurred during the recognition process with the SAs, which contributed to the surprisingly uniform affinity for all the SAs tested. This study verified the previous hypotheses on antibody production against SAs and enhanced our understanding of antibody-SAs interactions, which provided useful information toward the rational design of novel haptens and directed evolution to produce class-specific antibodies as DHPS.
- Beijing Advanced Innovation Center for Food Nutrition and Human Health, College of Veterinary Medicine , China Agricultural University , 100193 Beijing , People's Republic of China.
Organizational Affiliation: