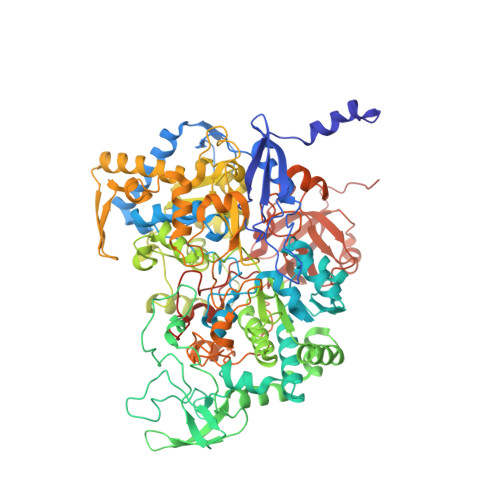
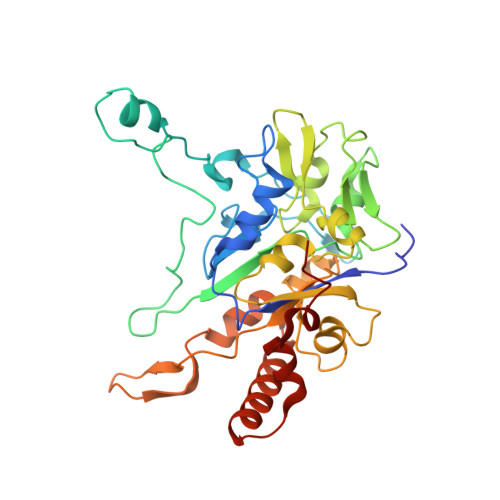
Perchlorate Reductase Is Distinguished by Active Site Aromatic Gate Residues.
Youngblut, M.D., Tsai, C.L., Clark, I.C., Carlson, H.K., Maglaqui, A.P., Gau-Pan, P.S., Redford, S.A., Wong, A., Tainer, J.A., Coates, J.D.(2016) J Biological Chem 291: 9190-9202
- PubMed: 26940877 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M116.714618
- Primary Citation Related Structures:
4YDD, 5CH7, 5CHC, 5E7O - PubMed Abstract:
Perchlorate is an important ion on both Earth and Mars. Perchlorate reductase (PcrAB), a specialized member of the dimethylsulfoxide reductase superfamily, catalyzes the first step of microbial perchlorate respiration, but little is known about the biochemistry, specificity, structure, and mechanism of PcrAB. Here we characterize the biophysics and phylogeny of this enzyme and report the 1.86-Å resolution PcrAB complex crystal structure. Biochemical analysis revealed a relatively high perchlorate affinity (Km = 6 μm) and a characteristic substrate inhibition compared with the highly similar respiratory nitrate reductase NarGHI, which has a relatively much lower affinity for perchlorate (Km = 1.1 mm) and no substrate inhibition. Structural analysis of oxidized and reduced PcrAB with and without the substrate analog SeO3 (2-) bound to the active site identified key residues in the positively charged and funnel-shaped substrate access tunnel that gated substrate entrance and product release while trapping transiently produced chlorate. The structures suggest gating was associated with shifts of a Phe residue between open and closed conformations plus an Asp residue carboxylate shift between monodentate and bidentate coordination to the active site molybdenum atom. Taken together, structural and mutational analyses of gate residues suggest key roles of these gate residues for substrate entrance and product release. Our combined results provide the first detailed structural insight into the mechanism of biological perchlorate reduction, a critical component of the chlorine redox cycle on Earth.
- From the Energy Biosciences Institute and.
Organizational Affiliation: