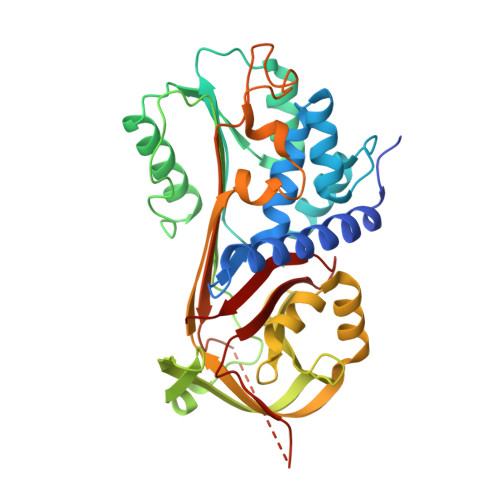
Smoothing a rugged protein folding landscape by sequence-based redesign.
Porebski, B.T., Keleher, S., Hollins, J.J., Nickson, A.A., Marijanovic, E.M., Borg, N.A., Costa, M.G., Pearce, M.A., Dai, W., Zhu, L., Irving, J.A., Hoke, D.E., Kass, I., Whisstock, J.C., Bottomley, S.P., Webb, G.I., McGowan, S., Buckle, A.M.(2016) Sci Rep 6: 33958-33958
- PubMed: 27667094 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep33958
- Primary Citation Related Structures:
5CDX, 5CDZ, 5CE0 - PubMed Abstract:
The rugged folding landscapes of functional proteins puts them at risk of misfolding and aggregation. Serine protease inhibitors, or serpins, are paradigms for this delicate balance between function and misfolding. Serpins exist in a metastable state that undergoes a major conformational change in order to inhibit proteases. However, conformational labiality of the native serpin fold renders them susceptible to misfolding, which underlies misfolding diseases such as α 1 -antitrypsin deficiency. To investigate how serpins balance function and folding, we used consensus design to create conserpin, a synthetic serpin that folds reversibly, is functional, thermostable, and polymerization resistant. Characterization of its structure, folding and dynamics suggest that consensus design has remodeled the folding landscape to reconcile competing requirements for stability and function. This approach may offer general benefits for engineering functional proteins that have risky folding landscapes, including the removal of aggregation-prone intermediates, and modifying scaffolds for use as protein therapeutics.
- Biomedicine Discovery Institute, Department of Biochemistry and Molecular Biology, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: