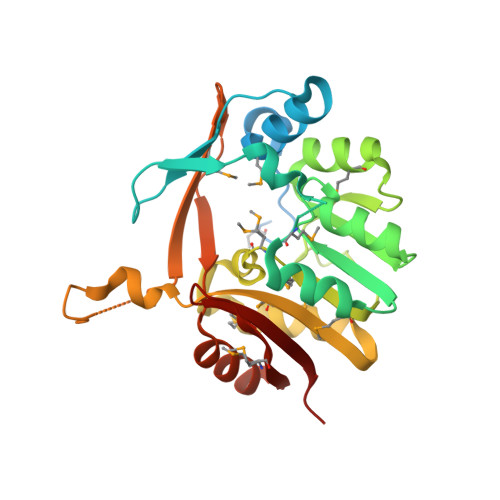
Crystal structure of SAM-dependent methyltransferase fromBacteroides fragilis in complex with S-Adenosyl-L-homocysteine
Gasiorowska, O.A., Shabalin, I.G., Handing, K.B., Cymborowski, M.T., Mason, D., Bonanno, J., Seidel, R., Almo, S.C., Minor, W.To be published.