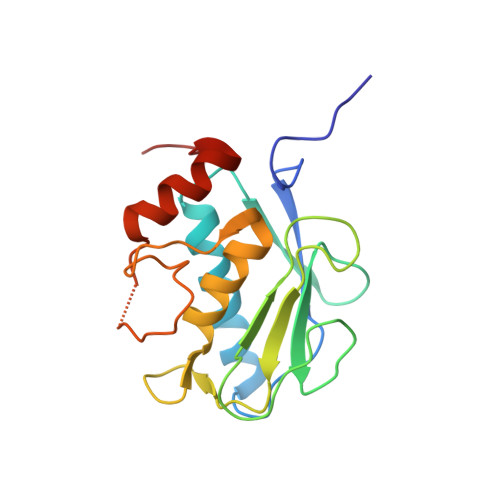
Fragment-based discovery of indole inhibitors of matrix metalloproteinase-13.
Taylor, S.J., Abeywardane, A., Liang, S., Muegge, I., Padyana, A.K., Xiong, Z., Hill-Drzewi, M., Farmer, B., Li, X., Collins, B., Li, J.X., Heim-Riether, A., Proudfoot, J., Zhang, Q., Goldberg, D., Zuvela-Jelaska, L., Zaher, H., Li, J., Farrow, N.A.(2011) J Med Chem 54: 8174-8187
- PubMed: 22017539 Search on PubMed
- DOI: https://doi.org/10.1021/jm201129m
- Primary Citation Related Structures:
5BOT, 5BOY, 5BPA - PubMed Abstract:
Matrix metalloproteases (MMPs) play an important role in cartilage homeostasis under both normal and inflamed disease states and, thus, have become attractive targets for the treatment of arthritic diseases. Herein, we describe the identification of a potent, selective MMP-13 inhibitor, developed using fragment-based structure-guided lead identification and optimization techniques. Virtual screening methods identified a novel, indole-based MMP-13 inhibitor that bound into the S1' pocket of the protein exhibiting a novel interaction pattern hitherto not observed in MMP-13 inhibitors. X-ray crystallographic structures were used to guide the elaboration of the fragment, ultimately leading to a potent inhibitor that was >100-fold selective over nine other MMP isoforms tested.
- Boehringer Ingelheim Pharmaceuticals Inc, Ridgefield, Connecticut 06877-0368, United States. steven.taylor@boehringeringelheim.com
Organizational Affiliation: