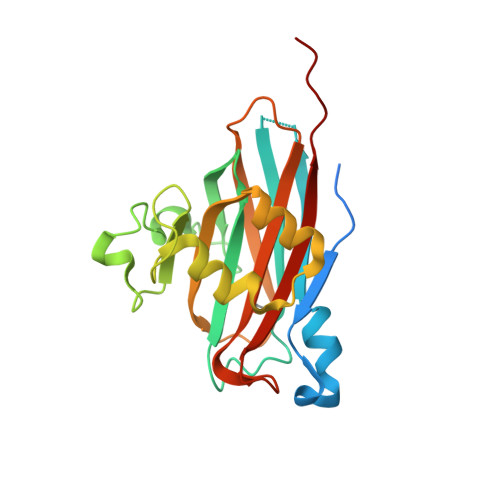
Expression, purification, crystallization and structure determination of the N terminal domain of Fhb, a factor H binding protein from Streptococcus suis.
Zhang, C., Yu, Y., Yang, M., Jiang, Y.(2015) Biochem Biophys Res Commun 466: 413-417
- PubMed: 26365348 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2015.09.040
- Primary Citation Related Structures:
5BOB - PubMed Abstract:
Fhb is a surface virulence protein from Streptococcus suis, which could aid bacterial evasion of host innate immune defense by recruiting complement regulator factor H to inactivate C3b deposited on bacterial surface in blood. Here we successfully expressed and purified the N terminal domain of Fhb (N-Fhb) and obtained crystals of the N-Fhb by sitting-drop vapor diffusion method with a resolution of 1.50 Å. The crystals belong to space group C2 with unit cell parameters a = 127.1 Å, b = 77.3 Å, c = 131.6 Å, α = 90°, β = 115.9°, γ = 90°. The structure of N-Fhb was determined by SAD method and the core structure of N-Fhb is a β sandwich. We speculated that binding of Fhb to human factor H may be mainly mediated by surface amino acids with negative charges.
- State Key Laboratory of Pathogen and Biosecurity, Beijng Institute of Microbiology and Infectious Disease, No. 20 Dongda Street, Fengtai District, Beijing 100071, China.
Organizational Affiliation: