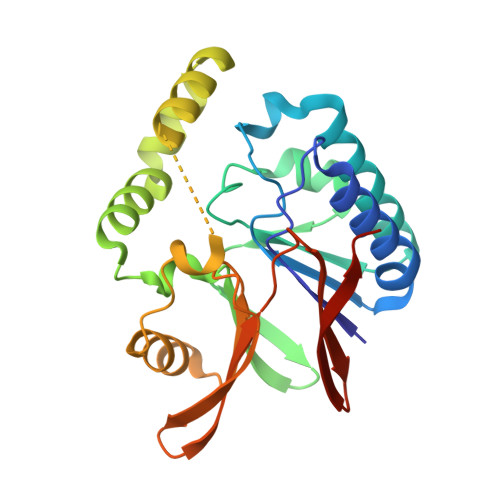
Crystal structures of the UDP-diacylglucosamine pyrophosphohydrase LpxH from Pseudomonas aeruginosa
Okada, C., Wakabayashi, H., Kobayashi, M., Shinoda, A., Tanaka, I., Yao, M.(2016) Sci Rep 6: 32822-32822
- PubMed: 27609419 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep32822
- Primary Citation Related Structures:
5B49, 5B4A, 5B4B, 5B4C, 5B4D - PubMed Abstract:
Lipid A (also known as endotoxin) is the hydrophobic portion of lipopolysaccharides. It is an essential membrane component required for the viability of gram-negative bacteria. The enzymes involved in its biosynthesis are attractive targets for the development of novel antibiotics. LpxH catalyzes the fourth step of the lipid A biosynthesis pathway and cleaves the pyrophosphate bond of UDP-2,3-diacylglucosamine to yield 2,3-diacylglucosamine 1-phosphate (lipid X) and UMP. Here we present the structures of LpxH from Pseudomonas aeruginosa (PaLpxH). PaLpxH consists of two domains: a catalytic domain that is homologous to the metallophosphoesterases and a helical insertion domain. Lipid X was captured in the crevice between these two domains, with its phosphate group facing the dinuclear metal (Mn(2+)) center and two acyl chains buried in the hydrophobic cavity. The structures reveal that a large conformational change occurs at the lipid X binding site surface upon the binding/release of the product molecule. Based on these observations, we propose a novel model for lipid X embedding, which involves the scissor-like movement of helix α6, resulting in the release of lipid X into the lipid bilayer.
- Faculty of Advanced Life Science, Hokkaido University, Sapporo 060-0810, Japan.
Organizational Affiliation: