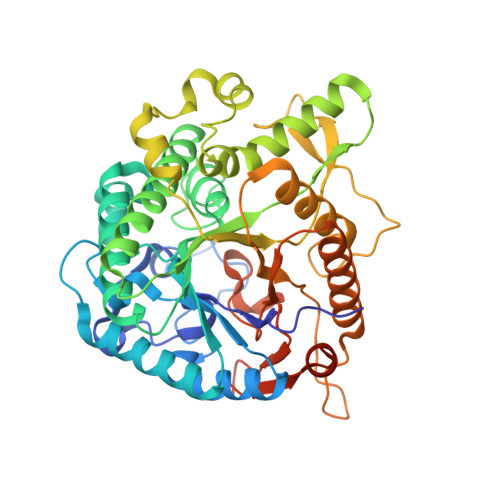
Crystal structure and identification of a key amino acid for glucose tolerance, substrate specificity, and transglycosylation activity of metagenomic beta-glucosidase Td2F2
Matsuzawa, T., Jo, T., Uchiyama, T., Manninen, J.A., Arakawa, T., Miyazaki, K., Fushinobu, S., Yaoi, K.(2016) FEBS J 283: 2340-2353
- PubMed: 27092463 Search on PubMed
- DOI: https://doi.org/10.1111/febs.13743
- Primary Citation Related Structures:
3WH5, 3WH6, 3WH7, 3WH8, 5AYB, 5AYI - PubMed Abstract:
β-Glucosidase Td2F2 isolated from a compost metagenome has high glucose tolerance and transglycosylation activity. In this study, we determined the high-resolution crystal structure of Td2F2. It has a unique structure at the -1 subsite that is important for substrate specificity but not for glucose tolerance. To elucidate the mechanism(s) of glucose tolerance, we isolated a glucose-sensitive Td2F2 mutant using random mutagenesis. In this mutant, Asn223 residue located between subsites +1 and +2 was mutated. The Asn223 mutation resulted in reduced glucose tolerance and transglycosylation activity, and drastically changed substrate specificity. These results indicate that the structure between subsites +1 and +2 is critical for the glucose tolerance and substrate specificity of Td2F2. Our findings shed light on the glucose tolerance and transglycosylation activity mechanisms of glycoside hydrolase family 1 β-glucosidases. The atomic coordinates and structure factors (codes 3WH5, 3WH6, 3WH8, 3WH7, 5AYB, and 5AYI) have been deposited in the Protein Data Bank (http://wwpdb.org/).
- Bioproduction Research Institute, National Institute of Advanced Industrial Science and Technology (AIST), Tsukuba, Ibaraki, Japan.
Organizational Affiliation: