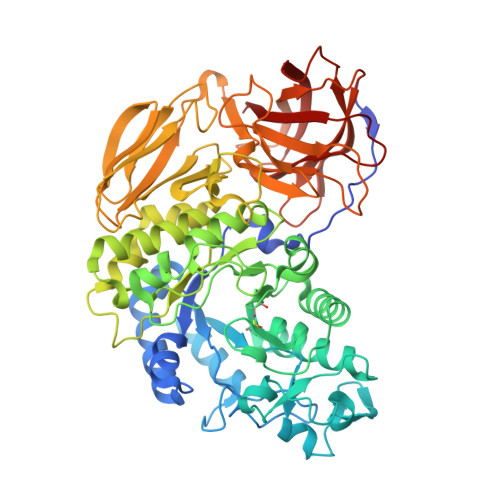
Crystal Structure and Mutational Analysis of Isomalto-dextranase, a Member of Glycoside Hydrolase Family 27
Okazawa, Y., Miyazaki, T., Yokoi, G., Ishizaki, Y., Nishikawa, A., Tonozuka, T.(2015) J Biological Chem 290: 26339-26349
- PubMed: 26330557 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.680942
- Primary Citation Related Structures:
5AWO, 5AWP, 5AWQ - PubMed Abstract:
Arthrobacter globiformis T6 isomalto-dextranase (AgIMD) is an enzyme that liberates isomaltose from the non-reducing end of a polymer of glucose, dextran. AgIMD is classified as a member of the glycoside hydrolase family (GH) 27, which comprises mainly α-galactosidases and α-N-acetylgalactosaminidases, whereas AgIMD does not show α-galactosidase or α-N-acetylgalactosaminidase activities. Here, we determined the crystal structure of AgIMD. AgIMD consists of the following three domains: A, C, and D. Domains A and C are identified as a (β/α)8-barrel catalytic domain and an antiparallel β-structure, respectively, both of which are commonly found in GH27 enzymes. However, domain A of AgIMD has subdomain B, loop-1, and loop-2, all of which are not found in GH27 human α-galactosidase. AgIMD in a complex with trisaccharide panose shows that Asp-207, a residue in loop-1, is involved in subsite +1. Kinetic parameters of the wild-type and mutant enzymes for the small synthetic saccharide p-nitrophenyl α-isomaltoside and the polysaccharide dextran were compared, showing that Asp-207 is important for the catalysis of dextran. Domain D is classified as carbohydrate-binding module (CBM) 35, and an isomaltose molecule is seen in this domain in the AgIMD-isomaltose complex. Domain D is highly homologous to CBM35 domains found in GH31 and GH66 enzymes. The results here indicate that some features found in GH13, -31, and -66 enzymes, such as subdomain B, residues at the subsite +1, and the CBM35 domain, are also observed in the GH27 enzyme AgIMD and thus provide insights into the evolutionary relationships among GH13, -27, -31, -36, and -66 enzymes.
- From the Department of Applied Biological Science, Tokyo University of Agriculture and Technology, Tokyo 183-8509, Japan.
Organizational Affiliation: