Multiple Capsid-Stabilizing Interactions Revealed in a High-Resolution Structure of an Emerging Picornavirus Causing Neonatal Sepsis
Shakeel, S., Westerhuis, B.M., Domanska, A., Koning, R.I., Matadeen, R., Koster, A.J., Bakker, A.Q., Beaumont, T., Wolthers, K.C., Butcher, S.J.(2016) Nat Commun 7: 11387
- PubMed: 27435188 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms11387
- Primary Citation Related Structures:
5APM - PubMed Abstract:
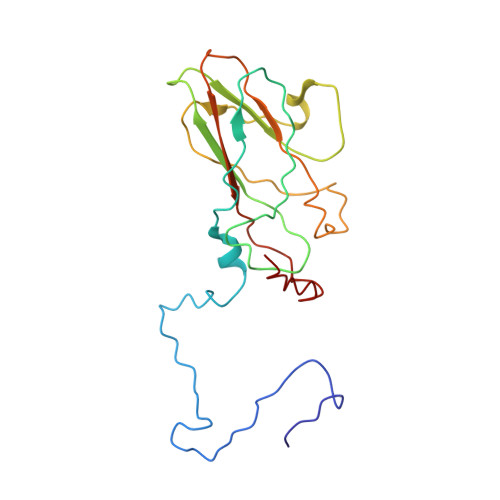
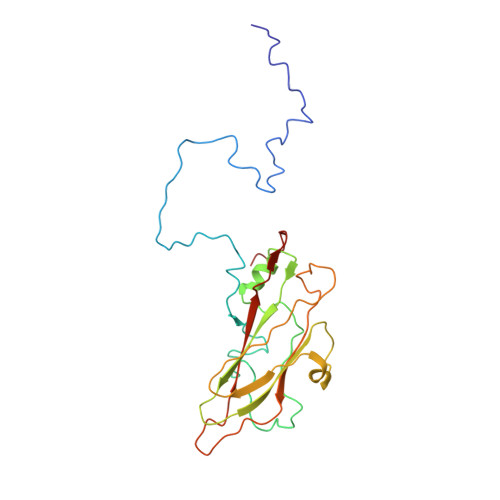
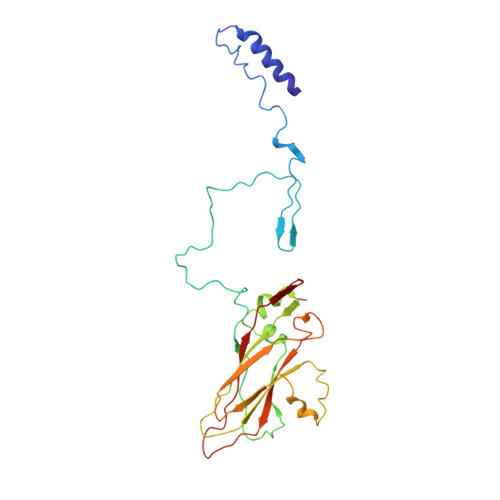
The poorly studied picornavirus, human parechovirus 3 (HPeV3) causes neonatal sepsis with no therapies available. Our 4.3-Å resolution structure of HPeV3 on its own and at 15 Å resolution in complex with human monoclonal antibody Fabs demonstrates the expected picornavirus capsid structure with three distinct features. First, 25% of the HPeV3 RNA genome in 60 sites is highly ordered as confirmed by asymmetric reconstruction, and interacts with conserved regions of the capsid proteins VP1 and VP3. Second, the VP0 N terminus stabilizes the capsid inner surface, in contrast to other picornaviruses where on expulsion as VP4, it forms an RNA translocation channel. Last, VP1's hydrophobic pocket, the binding site for the antipicornaviral drug, pleconaril, is blocked and thus inappropriate for antiviral development. Together, these results suggest a direction for development of neutralizing antibodies, antiviral drugs based on targeting the RNA-protein interactions and dissection of virus assembly on the basis of RNA nucleation.
- Institute of Biotechnology and Department of Biological Sciences, Viikinkaari 1, 00014 University of Helsinki, Finland.
Organizational Affiliation: