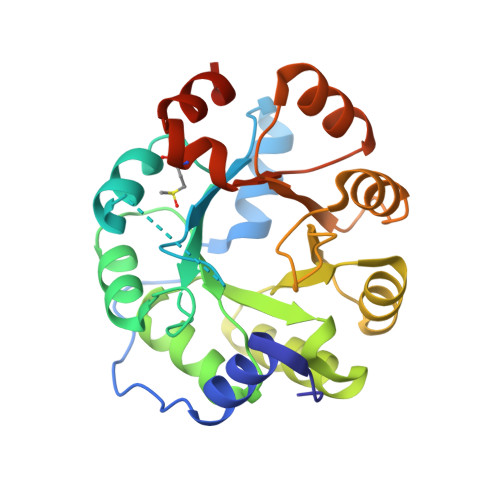
Emergence of a catalytic tetrad during evolution of a highly active artificial aldolase.
Obexer, R., Godina, A., Garrabou, X., Mittl, P.R., Baker, D., Griffiths, A.D., Hilvert, D.(2017) Nat Chem 9: 50-56
- PubMed: 27995916 Search on PubMed
- DOI: https://doi.org/10.1038/nchem.2596
- Primary Citation Related Structures:
5AN7, 5AOU - PubMed Abstract:
Designing catalysts that achieve the rates and selectivities of natural enzymes is a long-standing goal in protein chemistry. Here, we show that an ultrahigh-throughput droplet-based microfluidic screening platform can be used to improve a previously optimized artificial aldolase by an additional factor of 30 to give a >10 9 rate enhancement that rivals the efficiency of class I aldolases. The resulting enzyme catalyses a reversible aldol reaction with high stereoselectivity and tolerates a broad range of substrates. Biochemical and structural studies show that catalysis depends on a Lys-Tyr-Asn-Tyr tetrad that emerged adjacent to a computationally designed hydrophobic pocket during directed evolution. This constellation of residues is poised to activate the substrate by Schiff base formation, promote mechanistically important proton transfers and stabilize multiple transition states along a complex reaction coordinate. The emergence of such a sophisticated catalytic centre shows that there is nothing magical about the catalytic activities or mechanisms of naturally occurring enzymes, or the evolutionary process that gave rise to them.
- Laboratory of Organic Chemistry, ETH Zürich, 8093 Zürich, Switzerland.
Organizational Affiliation: