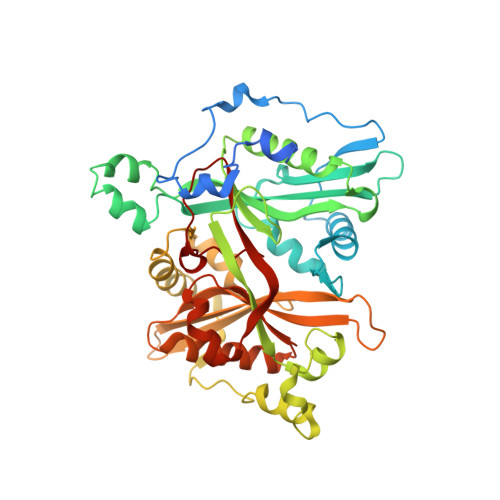
Development of Small-Molecule Trypanosoma Brucei N-Myristoyltransferase Inhibitors: Discovery and Optimisation of a Novel Binding Mode.
Spinks, D., Smith, V., Thompson, S., Robinson, D.A., Luksch, T., Smith, A., Torrie, L.S., Mcelroy, S., Stojanovski, L., Norval, S., Collie, I.T., Hallyburton, I., Rao, B., Brand, S., Brenk, R., Frearson, J.A., Read, K.D., Wyatt, P.G., Gilbert, I.H.(2015) ChemMedChem 10: 1821
- PubMed: 26395087 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cmdc.201500301
- Primary Citation Related Structures:
5AG4, 5AG5, 5AG6, 5AG7, 5AGE - PubMed Abstract:
The enzyme N-myristoyltransferase (NMT) from Trypanosoma brucei has been validated both chemically and biologically as a potential drug target for human African trypanosomiasis. We previously reported the development of some very potent compounds based around a pyrazole sulfonamide series, derived from a high-throughput screen. Herein we describe work around thiazolidinone and benzomorpholine scaffolds that were also identified in the screen. An X-ray crystal structure of the thiazolidinone hit in Leishmania major NMT showed the compound bound in the previously reported active site, utilising a novel binding mode. This provides potential for further optimisation. The benzomorpholinone was also found to bind in a similar region. Using an X-ray crystallography/structure-based design approach, the benzomorpholinone series was further optimised, increasing activity against T. brucei NMT by >1000-fold. A series of trypanocidal compounds were identified with suitable in vitro DMPK properties, including CNS exposure for further development. Further work is required to increase selectivity over the human NMT isoform and activity against T. brucei.
- Drug Discovery Unit, College of Life Sciences, University of Dundee, Sir James Black Centre, Dundee, DD1 5EH, UK.
Organizational Affiliation: