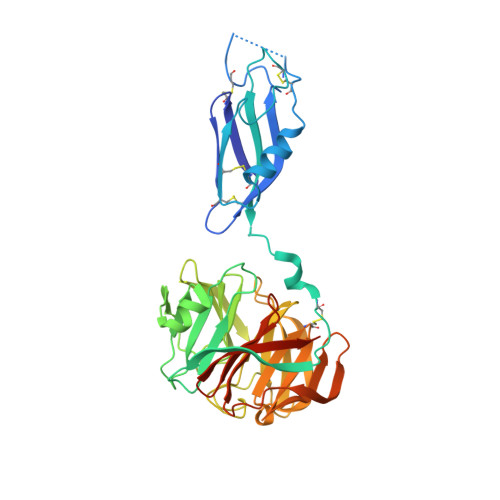
Structural Basis of Latrophilin-Flrt Interaction.
Jackson, V.A., Del Toro, D., Carrasquero, M., Roversi, P., Harlos, K., Klein, R., Seiradake, E.(2015) Structure 23: 774
- PubMed: 25728924 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.01.013
- Primary Citation Related Structures:
5AFB - PubMed Abstract:
Latrophilins, receptors for spider venom α-latrotoxin, are adhesion type G-protein-coupled receptors with emerging functions in synapse development. The N-terminal region binds the endogenous cell adhesion molecule FLRT, a major regulator of cortical and synapse development. We present crystallographic data for the mouse Latrophilin3 lectin and olfactomedin-like (Olf) domains, thereby revealing the Olf β-propeller fold and conserved calcium-binding site. We locate the FLRT-Latrophilin binding surfaces by a combination of sequence conservation analysis, point mutagenesis, and surface plasmon resonance experiments. In stripe assays, we show that wild-type Latrophilin3 and its high-affinity interactor FLRT2, but not the binding-impaired mutants we generated, promote HeLa cell adhesion. In contrast, cortical neurons expressing endogenous FLRTs are repelled by wild-type Latrophilin3 and not by the binding-impaired mutant. Taken together, we present molecular level insights into Latrophilin structure, its FLRT-binding mechanism, and a role for Latrophilin and FLRT that goes beyond a simply adhesive interaction.
- Department of Biochemistry, Oxford University, South Parks Road, Oxford OX1 3QU, UK.
Organizational Affiliation: