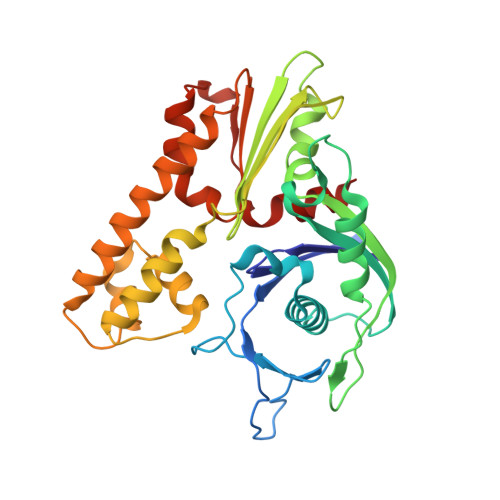
Structures of Actin-Like Parm Filaments Show Architecture of Plasmid-Segregating Spindles.
Bharat, T.A.M., Murshudov, G.N., Sachse, C., Lowe, J.(2015) Nature 523: 106
- PubMed: 25915019 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature14356
- Primary Citation Related Structures:
5AEY, 5AI7 - PubMed Abstract:
Active segregation of Escherichia coli low-copy-number plasmid R1 involves formation of a bipolar spindle made of left-handed double-helical actin-like ParM filaments. ParR links the filaments with centromeric parC plasmid DNA, while facilitating the addition of subunits to ParM filaments. Growing ParMRC spindles push sister plasmids to the cell poles. Here, using modern electron cryomicroscopy methods, we investigate the structures and arrangements of ParM filaments in vitro and in cells, revealing at near-atomic resolution how subunits and filaments come together to produce the simplest known mitotic machinery. To understand the mechanism of dynamic instability, we determine structures of ParM filaments in different nucleotide states. The structure of filaments bound to the ATP analogue AMPPNP is determined at 4.3 Å resolution and refined. The ParM filament structure shows strong longitudinal interfaces and weaker lateral interactions. Also using electron cryomicroscopy, we reconstruct ParM doublets forming antiparallel spindles. Finally, with whole-cell electron cryotomography, we show that doublets are abundant in bacterial cells containing low-copy-number plasmids with the ParMRC locus, leading to an asynchronous model of R1 plasmid segregation.
- Structural Studies Division, MRC Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge CB2 0QH, UK.
Organizational Affiliation: