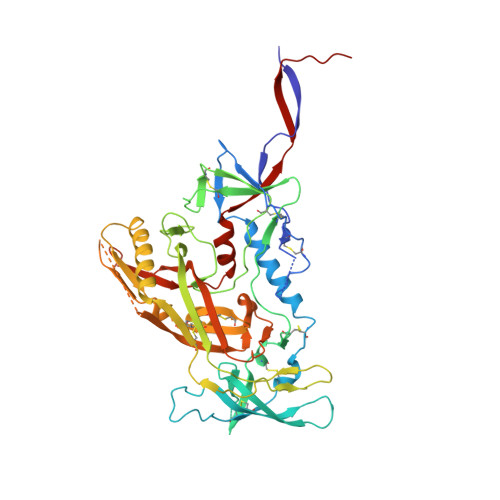
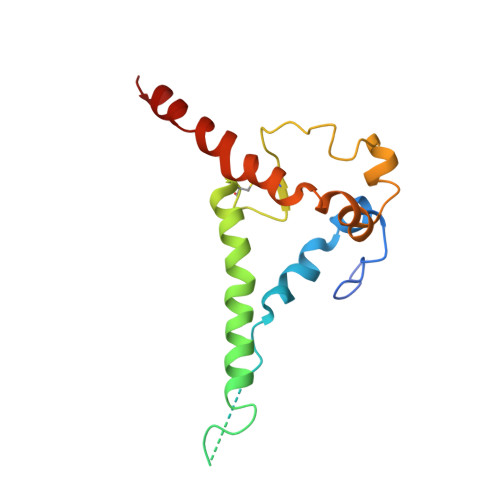
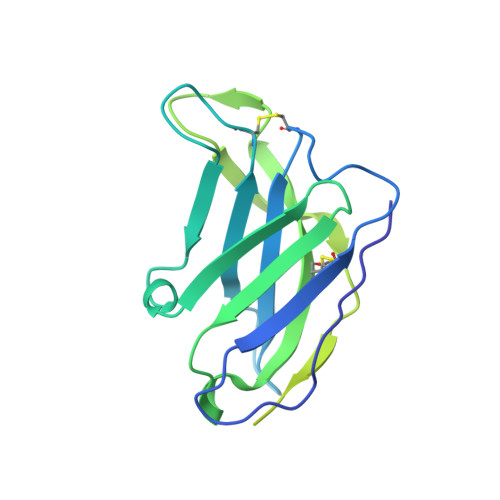
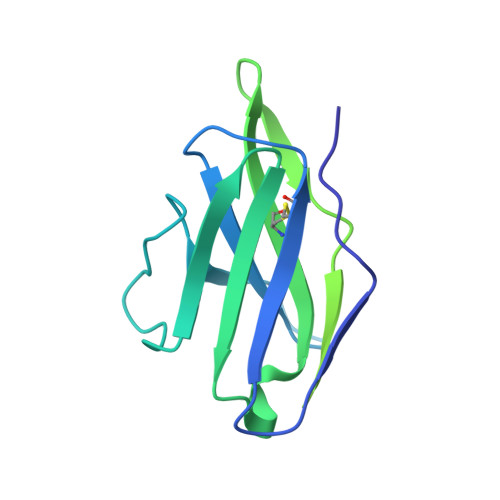
Model Building and Refinement of a Natively Glycosylated HIV-1 Env Protein by High-Resolution Cryoelectron Microscopy.
Lee, J.H., De Val, N., Lyumkis, D., Ward, A.B.(2015) Structure 23: 1943
- PubMed: 26388028 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.07.020
- Primary Citation Related Structures:
5ACO - PubMed Abstract:
Secretory and membrane proteins from mammalian cells undergo post-translational modifications, including N-linked glycosylation, which can result in a large number of possible glycoforms. This sample heterogeneity can be problematic for structural studies, particularly X-ray crystallography. Thus, crystal structures of heavily glycosylated proteins such as the HIV-1 Env viral spike protein have been determined by removing the majority of glycans. This step is most frequently carried out using Endoglycosidase H (EndoH) and requires that all expressed glycans be in the high-mannose form, which is often not the native glycoform. With significantly improved technologies in single-particle cryoelectron microscopy, we demonstrate that it is now possible to refine and build natively glycosylated HIV-1 Env structures in solution to 4.36 Å resolution. At this resolution we can now analyze the complete epitope of a broadly neutralizing antibody (bnAb), PGT128, in the context of the trimer expressed with native glycans.
- Department of Integrative Structural and Computational Biology, Center for HIV/AIDS Vaccine Immunology and Immunogen Discovery, The Scripps Research Institute, La Jolla, CA 92037, USA; International AIDS Vaccine Initiative (IAVI) Neutralizing Antibody Center and Collaboration for AIDS Vaccine Discovery (CAVD), The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: