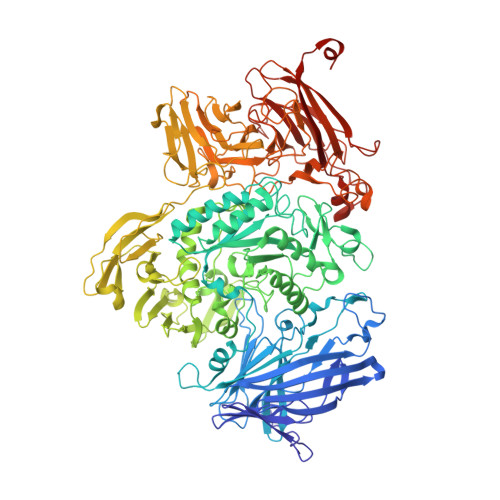
Structural Analysis of a Family 101 Glycoside Hydrolase in Complex with Carbohydrates Reveals Insights Into its Mechanism.
Gregg, K.J., Suits, M.D.L., Deng, L., Vocadlo, D.J., Boraston, A.B.(2015) J Biological Chem 290: 25657
- PubMed: 26304114 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.680470
- Primary Citation Related Structures:
5A55, 5A56, 5A57, 5A58, 5A59, 5A5A - PubMed Abstract:
O-Linked glycosylation is one of the most abundant post-translational modifications of proteins. Within the secretory pathway of higher eukaryotes, the core of these glycans is frequently an N-acetylgalactosamine residue that is α-linked to serine or threonine residues. Glycoside hydrolases in family 101 are presently the only known enzymes to be able to hydrolyze this glycosidic linkage. Here we determine the high-resolution structures of the catalytic domain comprising a fragment of GH101 from Streptococcus pneumoniae TIGR4, SpGH101, in the absence of carbohydrate, and in complex with reaction products, inhibitor, and substrate analogues. Upon substrate binding, a tryptophan lid (residues 724-WNW-726) closes on the substrate. The closing of this lid fully engages the substrate in the active site with Asp-764 positioned directly beneath C1 of the sugar residue bound within the -1 subsite, consistent with its proposed role as the catalytic nucleophile. In all of the bound forms of the enzyme, however, the proposed catalytic acid/base residue was found to be too distant from the glycosidic oxygen (>4.3 Å) to serve directly as a general catalytic acid/base residue and thereby facilitate cleavage of the glycosidic bond. These same complexes, however, revealed a structurally conserved water molecule positioned between the catalytic acid/base and the glycosidic oxygen. On the basis of these structural observations we propose a new variation of the retaining glycoside hydrolase mechanism wherein the intervening water molecule enables a Grotthuss proton shuttle between Glu-796 and the glycosidic oxygen, permitting this residue to serve as the general acid/base catalytic residue.
- From the Department of Biochemistry and Microbiology, University of Victoria, Victoria, British Columbia, V8W 3P6 and.
Organizational Affiliation: