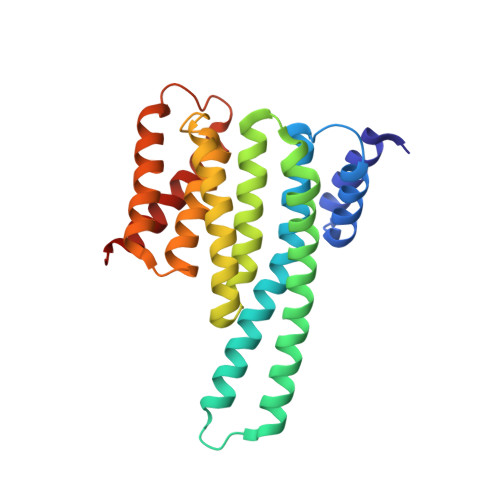
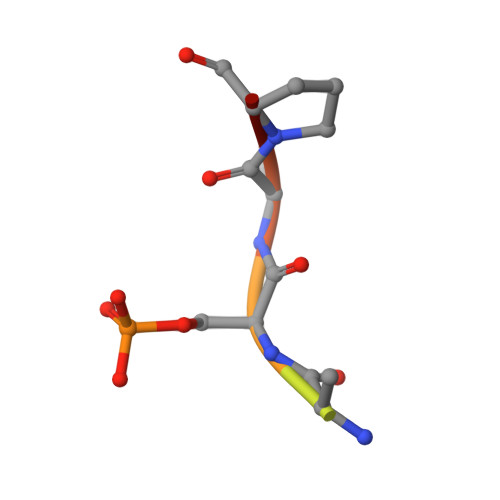
Molecular Dynamics Simulations and Structural Analysis of Giardia duodenalis 14-3-3 Protein-Protein Interactions.
Cau, Y., Fiorillo, A., Mori, M., Ilari, A., Botta, M., Lalle, M.(2015) J Chem Inf Model 55: 2611-2622
- PubMed: 26551337 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jcim.5b00452
- Primary Citation Related Structures:
4ZQ0, 5BY9 - PubMed Abstract:
Giardiasis is a gastrointestinal diarrheal illness caused by the protozoan parasite Giardia duodenalis, which affects annually over 200 million people worldwide. The limited antigiardial drug arsenal and the emergence of clinical cases refractory to standard treatments dictate the need for new chemotherapeutics. The 14-3-3 family of regulatory proteins, extensively involved in protein-protein interactions (PPIs) with pSer/pThr clients, represents a highly promising target. Despite homology with human counterparts, the single 14-3-3 of G. duodenalis (g14-3-3) is characterized by a constitutive phosphorylation in a region critical for target binding, thus affecting the function and the conformation of g14-3-3/clients interaction. However, to approach the design of specific small molecule modulators of g14-3-3 PPIs, structural elucidations are required. Here, we present a detailed computational and crystallographic study exploring the implications of g14-3-3 phosphorylation on protein structure and target binding. Self-Guided Langevin Dynamics and classical molecular dynamics simulations show that phosphorylation affects locally and globally g14-3-3 conformation, inducing a structural rearrangement more suitable for target binding. Profitable features for g14-3-3/clients interaction were highlighted using a hydrophobicity-based descriptor to characterize g14-3-3 client peptides. Finally, the X-ray structure of g14-3-3 in complex with a mode-1 prototype phosphopeptide was solved and combined with structure-based simulations to identify molecular features relevant for clients binding to g14-3-3. The data presented herein provide a further and structural understanding of g14-3-3 features and set the basis for drug design studies.
- Department of Biotechnology, Chemistry and Pharmacy, University of Siena , via Aldo Moro 2, 53019 Siena, Italy.
Organizational Affiliation: