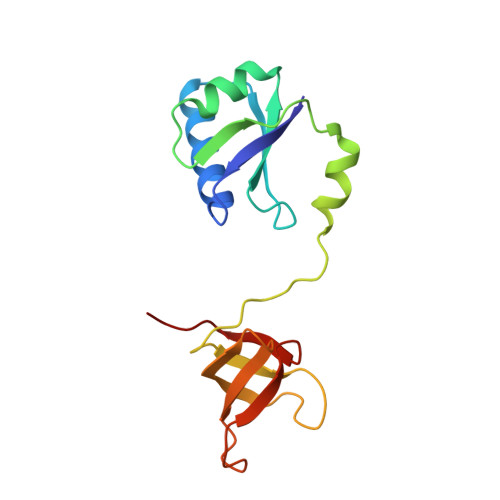
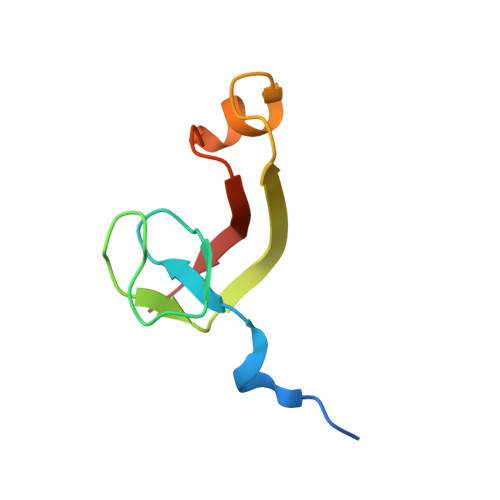
Structural and biochemical insights into the DNA-binding mode of MjSpt4p:Spt5 complex at the exit tunnel of RNAPII
Guo, G.R., Gao, Y.X., Zhu, Z.L., Zhao, D., Liu, Z., Zhou, H.H., Niu, L.W., Teng, M.K.(2015) J Struct Biol 192: 418-425
- PubMed: 26433031 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2015.09.023
- Primary Citation Related Structures:
4ZN1, 4ZN3 - PubMed Abstract:
Spt5 (NusG in bacteria) is the only RNA polymerase-associated factor known to be conserved in all three domains of life. In archaea and eukaryotes, Spt5 associates with Spt4, an elongation factor that is absent in bacteria, to form a functional heterodimeric complex. Previous studies suggest that the Spt4:Spt5 complex interacts directly with DNA at the double-stranded DNA exit tunnel of RNA polymerase to regulate gene transcription. In this study, the DNA-binding ability of Spt4:Spt5 from the archaeon Methanocaldococcus jannaschii was confirmed via nuclear magnetic resonance chemical shift perturbation and fluorescence polarization assays. Crystallographic analysis of the full-length MjSpt4:Spt5 revealed two distinct conformations of the C-terminal KOW domain of Spt5. A similar alkaline region was found on the Spt4:Spt5 surface in both crystal forms, and identified as double-stranded DNA binding patch through mutagenesis-fluorescence polarization assays. Based on these structural and biochemical data, the Spt4:Spt5-DNA binding model was built for the first time.
- Hefei National Laboratory for Physical Sciences at the Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230026, People's Republic of China; Key Laboratory of Structural Biology, Chinese Academy of Sciences, Hefei, Anhui 230026, People's Republic of China.
Organizational Affiliation: