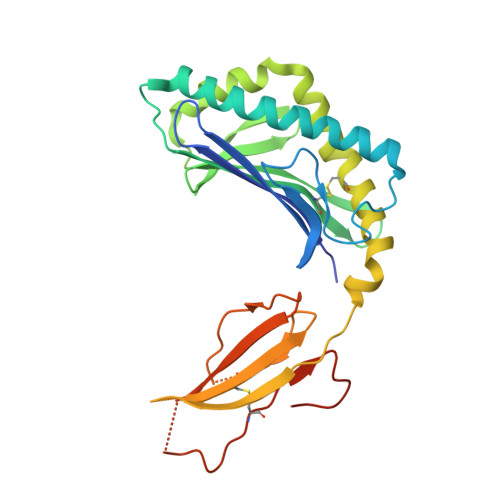
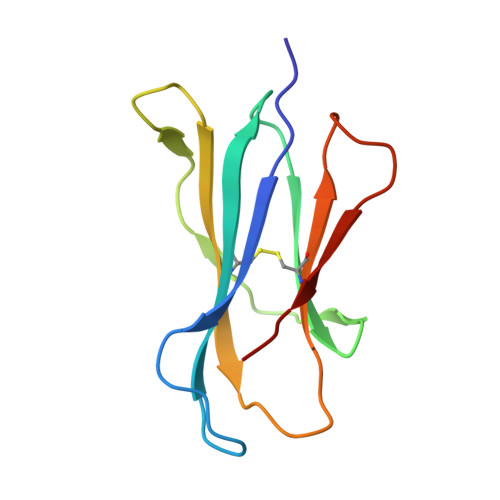
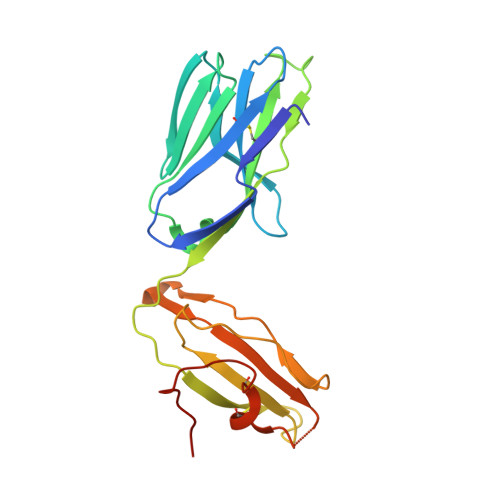
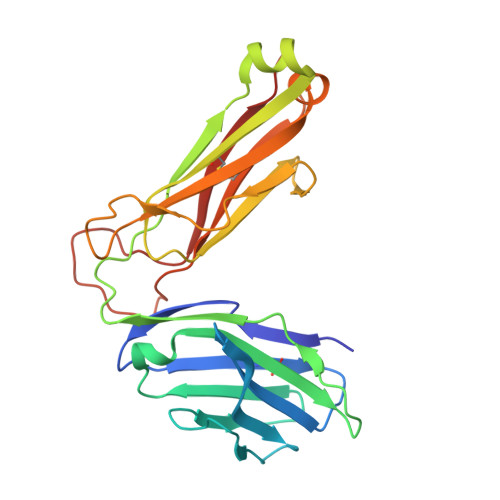
Structural modifications of alphaGalCer in both lipid and carbohydrate moiety influence activation of murine and human iNKT cells
Birkholz, A., Nemcovic, M., Yu, E.D., Girardi, E., Wang, J., Khurana, A., Pauwels, N., Franck, R.W., Tsuji, M., Howell, A., Calenbergh, S., Kronenberg, M., Zajonc, D.M.To be published.