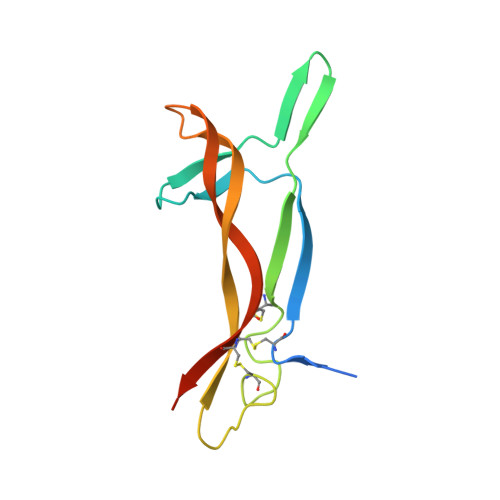
The structure of nerve growth factor in complex with lysophosphatidylinositol
Sun, H.L., Jiang, T.(2015) Acta Crystallogr F Struct Biol Commun 71: 906-912
- PubMed: 26144237 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X15008870
- Primary Citation Related Structures:
4XPJ - PubMed Abstract:
Nerve growth factor (NGF) is an important protein that is involved in a variety of physiological processes in cell survival, differentiation, proliferation and maintenance. The previously reported crystal structure of mouse NGF (mNGF) in complex with lysophosphatidylserine (LysoPS) showed that mNGF can bind LysoPS at its dimeric interface. To expand the understanding of the structural basis for specific lipid recognition by NGF, the crystal structure of mNGF complexed with lysophosphatidylinositol (13:0 LysoPI) was solved. Interestingly, in addition to Lys88, which interacts with the head glycerol group and the phosphate group of LysoPI, as seen in the mNGF-LysoPS structure, two additional residues, Tyr52 and Arg50, were found to assist in lipid binding by forming hydrogen bonds to the inositol moiety of the LysoPI molecule. The results suggest a specific recognition mechanism of inositol group-containing lipids by NGF, which may help in the design of bioactive compounds that can be delivered by NGF.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Beijing 100101, People's Republic of China.
Organizational Affiliation: