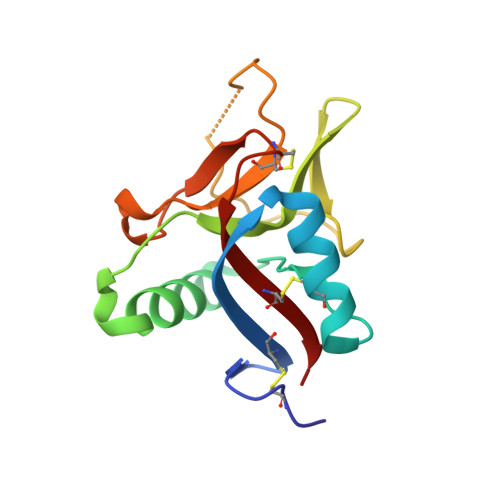
Crystal Structure of Struthiocalcin-1, an Intramineral Protein from Struthio Camelus Eggshell, in Two Different Crystal Forms.
Ruiz-Arellano, R.R., Medrano, F.J., Moreno, A., Romero, A.(2015) Acta Crystallogr D Biol Crystallogr 71: 809
- PubMed: 25849392 Search on PubMed
- DOI: https://doi.org/10.1107/S139900471500125X
- Primary Citation Related Structures:
4UWW, 4UXM - PubMed Abstract:
Biomineralization is the process by which living organisms produce minerals. One remarkable example is the formation of eggshells in birds. Struthiocalcins present in the ostrich (Struthio camellus) eggshell matrix act as biosensors of calcite growth during eggshell formation. Here, the crystal structure of struthiocalcin-1 (SCA-1) is reported in two different crystal forms. The structure is a compact single domain with an α/β fold characteristic of the C-type lectin family. In contrast to the related avian ovocleidin OC17, the electrostatic potential on the molecular surface is dominated by an acidic patch. Scanning electron microscopy combined with Raman spectroscopy indicates that these intramineral proteins (SCA-1 and SCA-2) induce calcium carbonate precipitation, leading to the formation of a stable form of calcite in the mature eggshell. Finally, the implications of these two intramineral proteins SCA-1 and SCA-2 in the nucleation of calcite during the formation of eggshells in ratite birds are discussed.
- Instituto de Química, Universidad Nacional Autónoma de México, 04510 Mexico City, Mexico.
Organizational Affiliation: