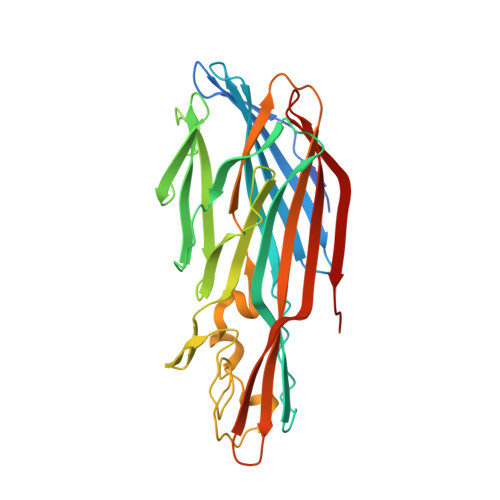
Crystal structures of the components of the Staphylococcus aureus leukotoxin ED.
Nocadello, S., Minasov, G., Shuvalova, L., Dubrovska, I., Sabini, E., Bagnoli, F., Grandi, G., Anderson, W.F.(2016) Acta Crystallogr D Struct Biol 72: 113-120
- PubMed: 26894539 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798315023207
- Primary Citation Related Structures:
3ROH, 4Q7G - PubMed Abstract:
Staphylococcal leukotoxins are a family of β-barrel, bicomponent, pore-forming toxins with membrane-damaging functions. These bacterial exotoxins share sequence and structural homology and target several host-cell types. Leukotoxin ED (LukED) is one of these bicomponent pore-forming toxins that Staphylococcus aureus produces in order to suppress the ability of the host to contain the infection. The recent delineation of the important role that LukED plays in S. aureus pathogenesis and the identification of its protein receptors, combined with its presence in S. aureus methicillin-resistant epidemic strains, establish this leukocidin as a possible target for the development of novel therapeutics. Here, the crystal structures of the water-soluble LukE and LukD components of LukED have been determined. The two structures illustrate the tertiary-structural variability with respect to the other leukotoxins while retaining the conservation of the residues involved in the interaction of the protomers in the bipartite leukotoxin in the pore complex.
- Center for Structural Genomics of Infectious Diseases, Department of Biochemistry and Molecular Genetics, Northwestern University Feinberg School of Medicine, Chicago, IL 60611, USA.
Organizational Affiliation: