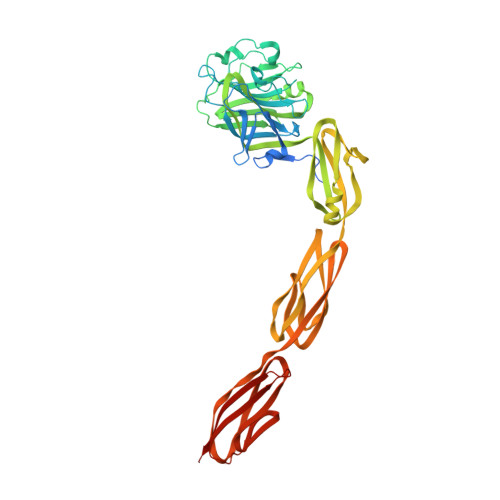
Structural Insights into SraP-Mediated Staphylococcus aureus Adhesion to Host Cells
Yang, Y.H., Jiang, Y.L., Zhang, J., Wang, L., Bai, X.H., Zhang, S.J., Ren, Y.M., Li, N., Zhang, Y.H., Zhang, Z., Gong, Q., Mei, Y., Xue, T., Zhang, J.R., Chen, Y., Zhou, C.Z.(2014) PLoS Pathog 10: e1004169-e1004169
- PubMed: 24901708 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1004169
- Primary Citation Related Structures:
4M00, 4M01, 4M02, 4M03 - PubMed Abstract:
Staphylococcus aureus, a Gram-positive bacterium causes a number of devastating human diseases, such as infective endocarditis, osteomyelitis, septic arthritis and sepsis. S. aureus SraP, a surface-exposed serine-rich repeat glycoprotein (SRRP), is required for the pathogenesis of human infective endocarditis via its ligand-binding region (BR) adhering to human platelets. It remains unclear how SraP interacts with human host. Here we report the 2.05 Å crystal structure of the BR of SraP, revealing an extended rod-like architecture of four discrete modules. The N-terminal legume lectin-like module specifically binds to N-acetylneuraminic acid. The second module adopts a β-grasp fold similar to Ig-binding proteins, whereas the last two tandem repetitive modules resemble eukaryotic cadherins but differ in calcium coordination pattern. Under the conditions tested, small-angle X-ray scattering and molecular dynamic simulation indicated that the three C-terminal modules function as a relatively rigid stem to extend the N-terminal lectin module outwards. Structure-guided mutagenesis analyses, in addition to a recently identified trisaccharide ligand of SraP, enabled us to elucidate that SraP binding to sialylated receptors promotes S. aureus adhesion to and invasion into host epithelial cells. Our findings have thus provided novel structural and functional insights into the SraP-mediated host-pathogen interaction of S. aureus.
- Hefei National Laboratory for Physical Sciences at the Microscale and School of Life Sciences, University of Science and Technology of China, Hefei Anhui, People's Republic of China.
Organizational Affiliation: