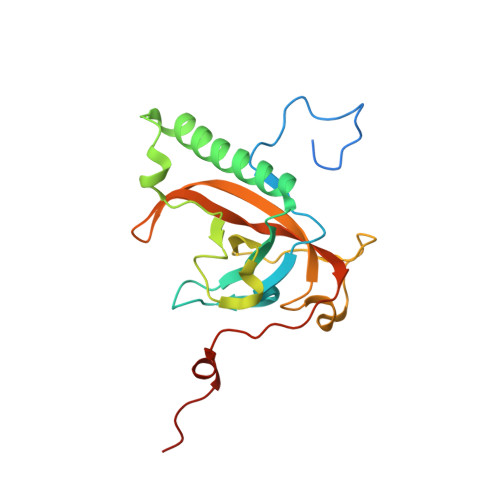
Structure of phage-related protein from Bacillus cereus ATCC 10987
Filippova, E.V., Wawrzak, Z., Minasov, G., Shuvalova, L., Kiryukhina, O., Babnigg, G., Rubin, E., Sacchettini, J., Joachimiak, A., Anderson, W.F., Midwest Center for Structural Genomics (MCSG), Structures of Mtb Proteins Conferring Susceptibility to Known Mtb Inhibitors (MTBI)To be published.