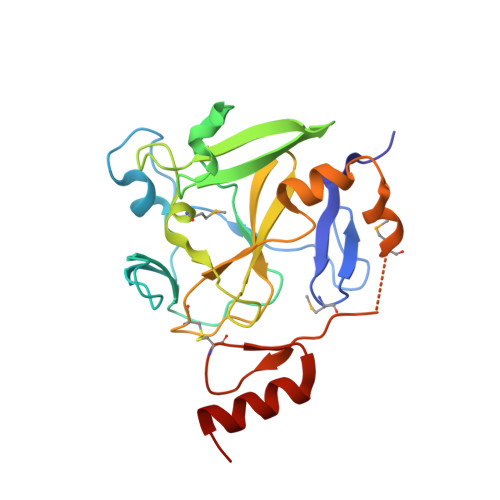
Crystal structure of methyltransferase domain of human PR domain-containing protein 9
Dombrovski, L., Dong, A., Li, Y., Tempel, W., Bountra, C., Arrowsmith, C.H., Edwards, A.M., Brown, P.J., Wu, H., Structural Genomics Consortium (SGC)To be published.