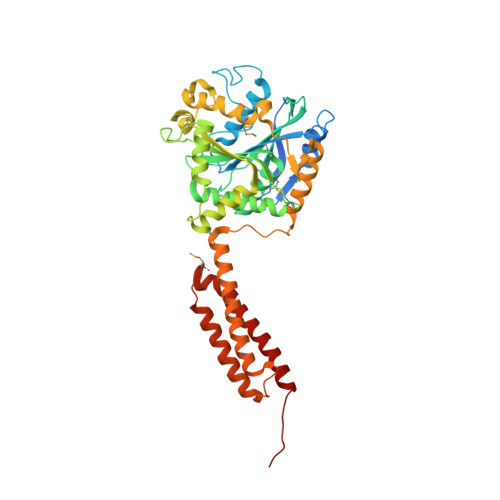
Structural basis for conformational switching and GTP loading of the large G protein atlastin.
Byrnes, L.J., Singh, A., Szeto, K., Benvin, N.M., O'Donnell, J.P., Zipfel, W.R., Sondermann, H.(2013) EMBO J 32: 369-384
- PubMed: 23334294 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2012.353
- Primary Citation Related Structures:
4IDN, 4IDO, 4IDP, 4IDQ - PubMed Abstract:
Atlastin, a member of the dynamin superfamily, is known to catalyse homotypic membrane fusion in the smooth endoplasmic reticulum (ER). Recent studies of atlastin have elucidated key features about its structure and function; however, several mechanistic details, including the catalytic mechanism and GTP hydrolysis-driven conformational changes, are yet to be determined. Here, we present the crystal structures of atlastin-1 bound to GDP·AlF(4)(-) and GppNHp, uncovering an intramolecular arginine finger that stimulates GTP hydrolysis when correctly oriented through rearrangements within the G domain. Utilizing Förster Resonance Energy Transfer, we describe nucleotide binding and hydrolysis-driven conformational changes in atlastin and their sequence. Furthermore, we discovered a nucleotide exchange mechanism that is intrinsic to atlastin's N-terminal domains. Our results indicate that the cytoplasmic domain of atlastin acts as a tether and homotypic interactions are timed by GTP binding and hydrolysis. Perturbation of these mechanisms may be implicated in a group of atlastin-associated hereditary neurodegenerative diseases.
- Department of Molecular Medicine, College of Veterinary Medicine, Cornell University, Ithaca, NY 14853, USA.
Organizational Affiliation: