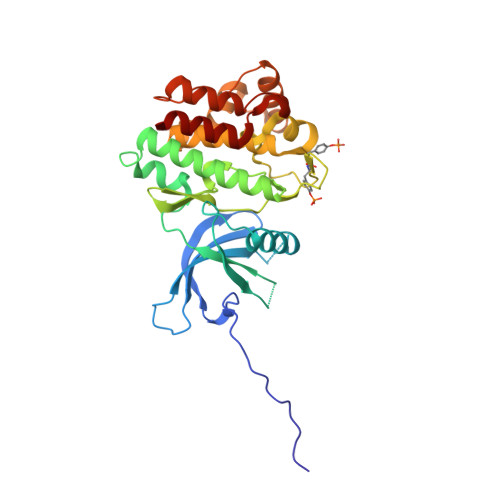
Identification of Imidazo-Pyrrolopyridines as Novel and Potent JAK1 Inhibitors.
Kulagowski, J.J., Blair, W., Bull, R.J., Chang, C., Deshmukh, G., Dyke, H.J., Eigenbrot, C., Ghilardi, N., Gibbons, P., Harrison, T.K., Hewitt, P.R., Liimatta, M., Hurley, C.A., Johnson, A., Johnson, T., Kenny, J.R., Bir Kohli, P., Maxey, R.J., Mendonca, R., Mortara, K., Murray, J., Narukulla, R., Shia, S., Steffek, M., Ubhayakar, S., Ultsch, M., van Abbema, A., Ward, S.I., Waszkowycz, B., Zak, M.(2012) J Med Chem 55: 5901-5921
- PubMed: 22591402 Search on PubMed
- DOI: https://doi.org/10.1021/jm300438j
- Primary Citation Related Structures:
4E4L, 4E4M, 4E4N, 4E5W, 4E6D, 4E6Q - PubMed Abstract:
A therapeutic rationale is proposed for the treatment of inflammatory diseases, such as rheumatoid arthritis (RA), by specific targeting of the JAK1 pathway. Examination of the preferred binding conformation of clinically effective, pan-JAK inhibitor 1 led to identification of a novel, tricyclic hinge binding scaffold 3. Exploration of SAR through a series of cycloamino and cycloalkylamino analogues demonstrated this template to be highly tolerant of substitution, with a predisposition to moderate selectivity for the JAK1 isoform over JAK2. This study culminated in the identification of subnanomolar JAK1 inhibitors such as 22 and 49, having excellent cell potency, good rat pharmacokinetic characteristics, and excellent kinase selectivity. Determination of the binding modes of the series in JAK1 and JAK2 by X-ray crystallography supported the design of analogues to enhance affinity and selectivity.
- Department of Medicinal Chemistry, Argenta, 8/9 Spire Green Centre, Harlow CM19 5TR, United Kingdom. janusz.kulagowski@glpg
Organizational Affiliation: