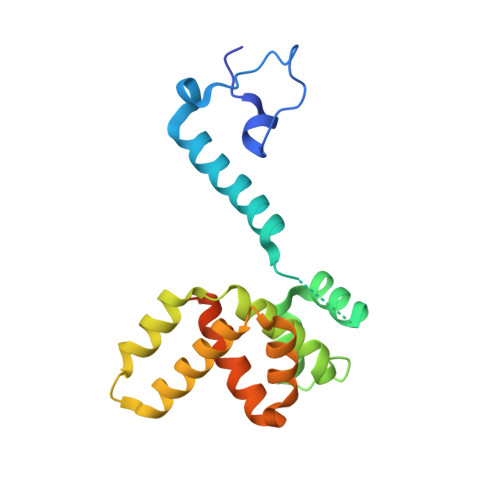
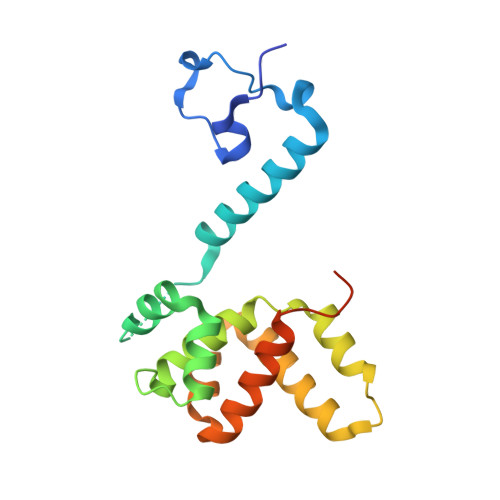
Drastic Changes in Conformational Dynamics of the Antiterminator M2-1 Regulate Transcription Efficiency in Pneumovirinae.
Leyrat, C., Renner, M., Harlos, K., Huiskonen, J.T., Grimes, J.M.(2014) Elife 3: 02674
- PubMed: 24842877 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.02674
- Primary Citation Related Structures:
4CS7, 4CS8, 4CS9, 4CSA - PubMed Abstract:
The M2-1 protein of human metapneumovirus (HMPV) is a zinc-binding transcription antiterminator which is highly conserved among pneumoviruses. We report the structure of tetrameric HMPV M2-1. Each protomer features a N-terminal zinc finger domain and an α-helical tetramerization motif forming a rigid unit, followed by a flexible linker and an α-helical core domain. The tetramer is asymmetric, three of the protomers exhibiting a closed conformation, and one an open conformation. Molecular dynamics simulations and SAXS demonstrate a dynamic equilibrium between open and closed conformations in solution. Structures of adenosine monophosphate- and DNA- bound M2-1 establish the role of the zinc finger domain in base-specific recognition of RNA. Binding to 'gene end' RNA sequences stabilized the closed conformation of M2-1 leading to a drastic shift in the conformational landscape of M2-1. We propose a model for recognition of gene end signals and discuss the implications of these findings for transcriptional regulation in pneumoviruses.DOI: http://dx.doi.org/10.7554/eLife.02674.001.
- Division of Structural Biology, Wellcome Trust Centre for Human Genetics, Oxford, United Kingdom.
Organizational Affiliation: