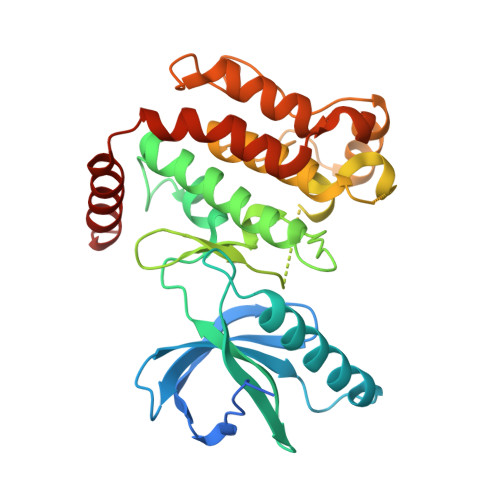
Inflammatory Signaling by NOD-RIPK2 Is Inhibited by Clinically Relevant Type II Kinase Inhibitors.
Canning, P., Ruan, Q., Schwerd, T., Hrdinka, M., Maki, J.L., Saleh, D., Suebsuwong, C., Ray, S., Brennan, P.E., Cuny, G.D., Uhlig, H.H., Gyrd-Hansen, M., Degterev, A., Bullock, A.N.(2015) Chem Biol 22: 1174-1184
- PubMed: 26320862 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2015.07.017
- Primary Citation Related Structures:
4C8B - PubMed Abstract:
RIPK2 mediates pro-inflammatory signaling from the bacterial sensors NOD1 and NOD2, and is an emerging therapeutic target in autoimmune and inflammatory diseases. We observed that cellular RIPK2 can be potently inhibited by type II inhibitors that displace the kinase activation segment, whereas ATP-competitive type I inhibition was only poorly effective. The most potent RIPK2 inhibitors were the US Food and Drug Administration-approved drugs ponatinib and regorafenib. Their mechanism of action was independent of NOD2 interaction and involved loss of downstream kinase activation as evidenced by lack of RIPK2 autophosphorylation. Notably, these molecules also blocked RIPK2 ubiquitination and, consequently, inflammatory nuclear factor κB signaling. In monocytes, the inhibitors selectively blocked NOD-dependent tumor necrosis factor production without affecting lipopolysaccharide-dependent pathways. We also determined the first crystal structure of RIPK2 bound to ponatinib, and identified an allosteric site for inhibitor development. These results highlight the potential for type II inhibitors to treat indications of RIPK2 activation as well as inflammation-associated cancers.
- Structural Genomics Consortium, University of Oxford, Old Road Campus, Roosevelt Drive, Oxford OX3 7DQ, UK.
Organizational Affiliation: