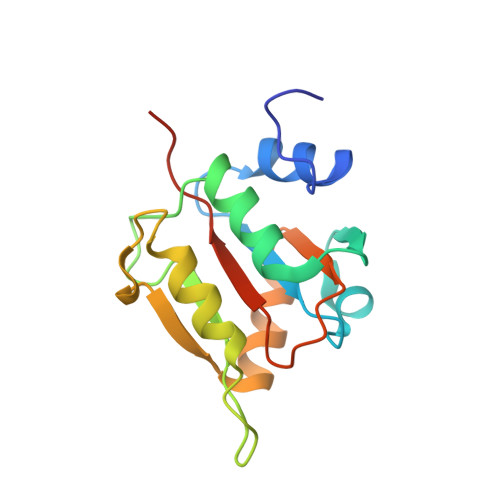
Structure of the Fucose Mutarotase from Streptococcus Pneumoniae in Complex with L-Fucose
Higgins, M.A., Boraston, A.B.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 1524
- PubMed: 22139157 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111046343
- Primary Citation Related Structures:
4A34 - PubMed Abstract:
Streptococcus pneumoniae relies on a variety of carbohydrate-utilization pathways for both colonization of its human host and full virulence during the development of invasive disease. One such pathway is the fucose-utilization pathway, a component of which is fucose mutarotase (SpFcsU), an enzyme that performs the interconversion between α-L-fucose and β-L-fucose. This protein was crystallized and its three-dimensional structure was solved in complex with L-fucose. The structure shows a complex decameric quaternary structure with a high overall degree of structural identity to Escherichia coli FcsU (EcFcsU). Furthermore, the active-site architecture of SpFcsU is highly similar to that of EcFcsU. When considered in the context of the fucose-utilization pathway found in S. pneumoniae, SpFcsU appears to link the two halves of the pathway by enhancing the rate of conversion of the product of the final glycoside hydrolysis step, β-fucose, into the substrate for the fucose isomerase, α-fucose.
- Department of Biochemistry and Microbiology, University of Victoria, Victoria, BC, Canada.
Organizational Affiliation: