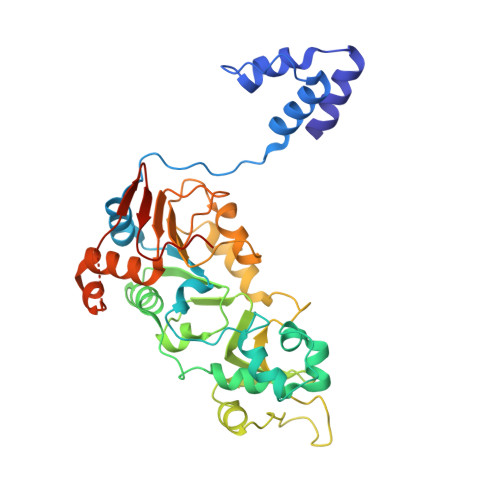
Crystallographic structure of a small molecule SIRT1 activator-enzyme complex.
Dai, H., Case, A.W., Riera, T.V., Considine, T., Lee, J.E., Hamuro, Y., Zhao, H., Jiang, Y., Sweitzer, S.M., Pietrak, B., Schwartz, B., Blum, C.A., Disch, J.S., Caldwell, R., Szczepankiewicz, B., Oalmann, C., Yee Ng, P., White, B.H., Casaubon, R., Narayan, R., Koppetsch, K., Bourbonais, F., Wu, B., Wang, J., Qian, D., Jiang, F., Mao, C., Wang, M., Hu, E., Wu, J.C., Perni, R.B., Vlasuk, G.P., Ellis, J.L.(2015) Nat Commun 6: 7645-7645
- PubMed: 26134520 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms8645
- Primary Citation Related Structures:
4ZZH, 4ZZI, 4ZZJ - PubMed Abstract:
SIRT1, the founding member of the mammalian family of seven NAD(+)-dependent sirtuins, is composed of 747 amino acids forming a catalytic domain and extended N- and C-terminal regions. We report the design and characterization of an engineered human SIRT1 construct (mini-hSIRT1) containing the minimal structural elements required for lysine deacetylation and catalytic activation by small molecule sirtuin-activating compounds (STACs). Using this construct, we solved the crystal structure of a mini-hSIRT1-STAC complex, which revealed the STAC-binding site within the N-terminal domain of hSIRT1. Together with hydrogen-deuterium exchange mass spectrometry (HDX-MS) and site-directed mutagenesis using full-length hSIRT1, these data establish a specific STAC-binding site and identify key intermolecular interactions with hSIRT1. The determination of the interface governing the binding of STACs with human SIRT1 facilitates greater understanding of STAC activation of this enzyme, which holds significant promise as a therapeutic target for multiple human diseases.
- 1] Sirtris, a GlaxoSmithKline Company, 200 Technology Square, Suite 300, Cambridge, Massachusetts 02139, USA [2] GlaxoSmithKline, 1250S. Collegeville Road, Collegeville, Pennsylvania 19426, USA.
Organizational Affiliation: